Course 02429 Analysis of correlated data: Mixed Linear Models Module 12: Repeated measures II, advanced methods Per Bruun Brockhoff
DTU Compute Building 324 - room 220 Technical University of Denmark 2800 Lyngby – Denmark e-mail: perbb@dtu.dk
Per Bruun Brockhoff (perbb@dtu.dk) Mixed Linear Models, Module 12 Fall 2014 1 / 24 Overview of this module
Overview of this module
1
Different view on the random effects approach Example: Activity of rats
2
Gaussian model of spatial correlation Example: Activity of rats
3
Other spatial correlation structures
4
Diagram of analysis
5
The semi-variogram
6
Analysing the time structure by polynomial regression
Per Bruun Brockhoff (perbb@dtu.dk) Mixed Linear Models, Module 12 Fall 2014 2 / 24 Aim of this module
Aim of this module
Extend the toolbox for dealing with repeated measures Introduce models where the covariance structure is directly specified Improve our ability to handle long series See how to specify these models in R
Per Bruun Brockhoff (perbb@dtu.dk) Mixed Linear Models, Module 12 Fall 2014 3 / 24 Aim of this module
Remember the rats data set?
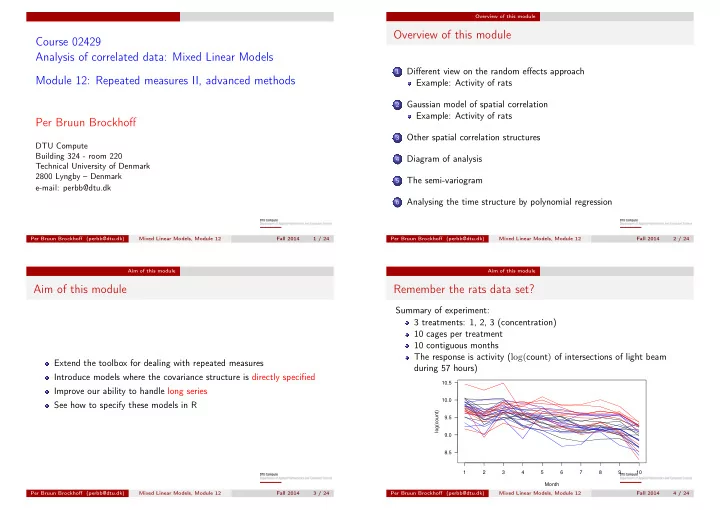
Summary of experiment: 3 treatments: 1, 2, 3 (concentration) 10 cages per treatment 10 contiguous months The response is activity (log(count) of intersections of light beam during 57 hours)
Month log(count) 8.5 9.0 9.5 10.0 10.5 1 2 3 4 5 6 7 8 9 10 Per Bruun Brockhoff (perbb@dtu.dk) Mixed Linear Models, Module 12 Fall 2014 4 / 24