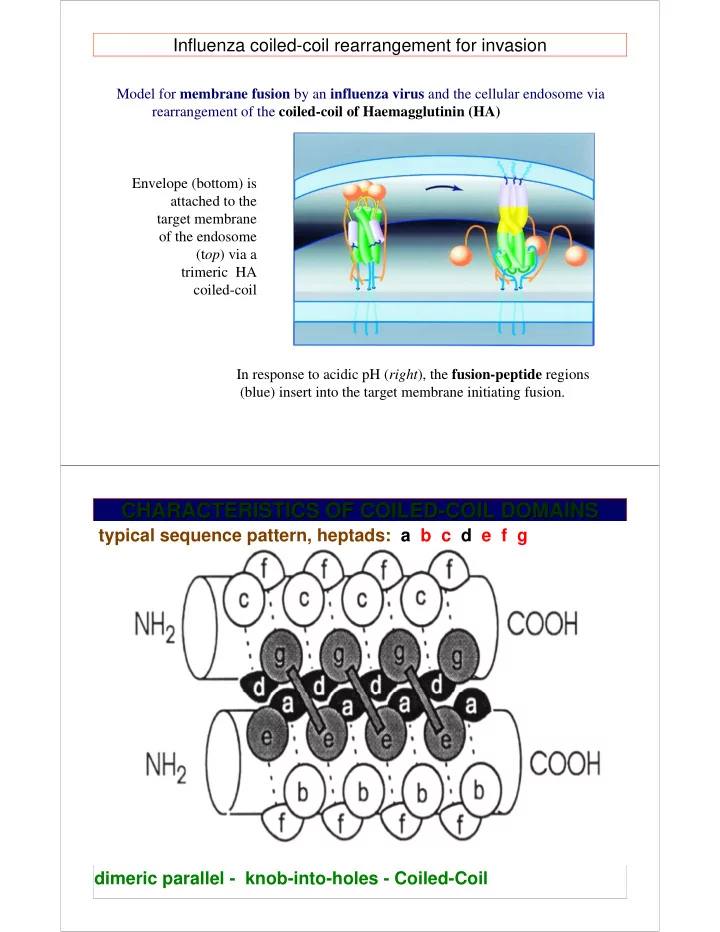
Model for membrane fusion by an influenza virus and the cellular endosome via rearrangement of the coiled-coil of Haemagglutinin (HA) In response to acidic pH (right), the fusion-peptide regions (blue) insert into the target membrane initiating fusion. Envelope (bottom) is attached to the target membrane
- f the endosome
(top) via a trimeric HA coiled-coil
Influenza coiled-coil rearrangement for invasion
CHARACTERISTICS OF COILED CHARACTERISTICS OF COILED-
- COIL DOMAINS
COIL DOMAINS
dimeric parallel - knob-into-holes - Coiled-Coil typical sequence pattern, heptads: a b c d e f g