Visualization, Summer Term 03 VIS, University of Stuttgart
1
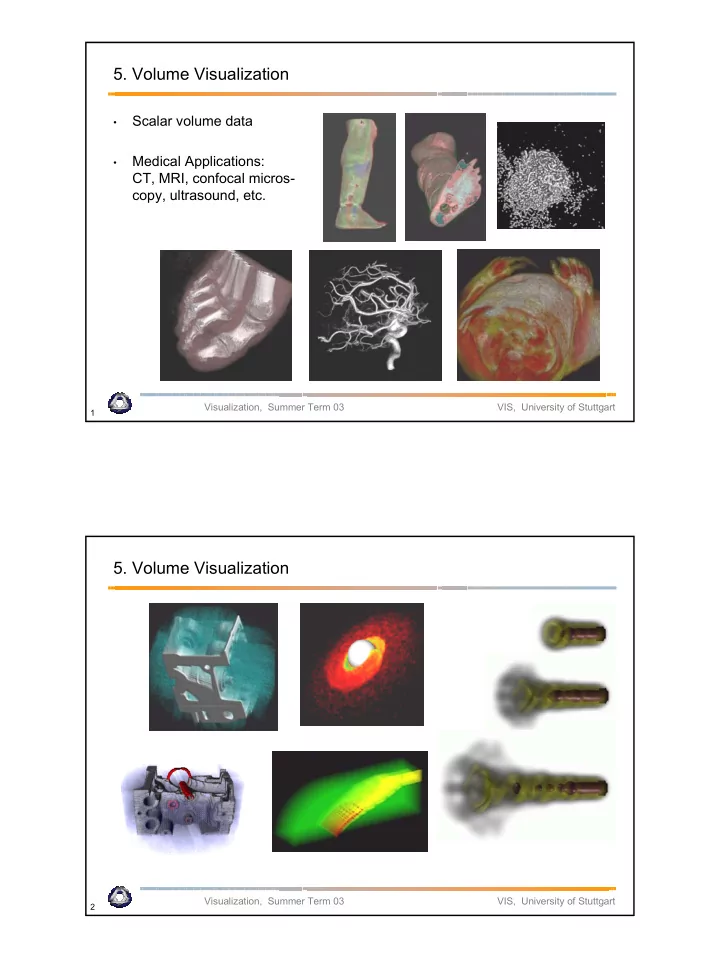
- 5. Volume Visualization
- Scalar volume data
- Medical Applications:
CT, MRI, confocal micros- copy, ultrasound, etc.
Visualization, Summer Term 03 VIS, University of Stuttgart
2
- 5. Volume Visualization