Theoretical Biology 2016 Transcription factors bind DNA to block - - PowerPoint PPT Presentation
Theoretical Biology 2016 Transcription factors bind DNA to block - - PowerPoint PPT Presentation
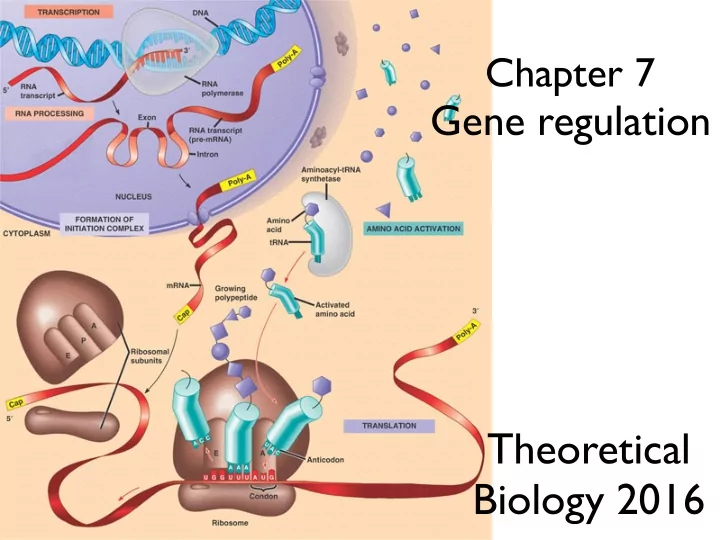
Chapter 7 Gene regulation Theoretical Biology 2016 Transcription factors bind DNA to block or enhance transcription From Campbell DNA makes RNA makes protein Golding et al. Cell 2005 d M d P d t = lM P d t = c dM and mRNA
Transcription factors
bind DNA to block or enhance transcription
From Campbell
DNA makes RNA makes protein
dM dt = c − dM and dP dt = lM − δP
Golding et al. Cell 2005
Golding et al. Cell 2005
mRNA formation occurs in bursts
Number of mRNA transcripts in individual cells over time. Red: data, blue: daughter cells, drops: cell division.
P M
c d
Mathematical model: mRNA & protein
dM dt = c − dM and dP dt = lM − δP Protein Messenger
Now with negative feedback on transcription
P M
c d
Messenger Protein
dM dt = c 1 + P/h − dM and dP dt = lM − δP ,
Quasi steady state assumption
Suppose turnover of protein much faster than that of mRNA
dP dt = lM − δP = 0
- r
P = l δ M
Substituting this into dM/dt gives:
dM dt = c 1 + (l/δ)M/h − dM = c 1 + M/h0 − dM
with h’ = hδ/l
dM dt = c 1 + P/h − dM and dP dt = lM − δP ,
Cross linking of receptors activates cells
allergen
Mast cell degranulation B cells activate and start to divide
Bivalent ligand binding a monovalent receptor
C: free ligand (C > NRT), R: free receptors, RT: total receptors, C1: single bound ligand, C2: double bound ligand: RT = R + C1 + 2C2 How does C2, and hence the growth rate, depend on C? C C2
dN dt = bN 2C2 RT − dN .
?
C C2 C1 R
R C C1 C2
RT = R + C1 + 2C2 , dC1 dt = 2konRC − koffC1 − xonRC1 + 2xoffC2 , dC2 dt = xonRC1 − 2xoffC2 . To study the steady state we set dC2/dt = 0 and add this to dC1/dt: dC1 dt = 0 = 2konRC − koffC1 = 2KRC − C1 , where K = kon/koff and R = RT − C1 − 2C2. Solving this gives C1 = 2CK(RT − 2C2) 1 + 2CK ,
10 0.3 0.15 C2 100 10 0.1 0.01 0.001 0.0001
- 05
1
C1 = 2CK(RT − 2C2) 1 + 2CK , which can be substituted into dC2/dt = 0 to solve C2 as a function of C: C2 = 1 + 4CK + 4C2K2 + 4CKRTX − (1 + 2CK)
q
(1 + 2CK)2 + 8CKRTX 8CKX where X = xon/xoff.
C2 C
Thus, the number of crosslinks is a bell- shaped function of the ligand concentration C. Cells grow best at intermediate ligand concentrations
dN dt = bN 2C2 RT − dN .
dt − − dC2 dt = xonRC1 − 2xoffC2 .
Lac operon, Jacob & Monod (1961)
From Campbell
Translate this into simple scheme
- peron: 0/1
repressor allolactose permease Β-galactos. Extra- cellular Lactose Cell membrane
Towards a phenomenological mathematical model
Repressor is modeled as a declining sigmoid Hill function. We will even scale the allolactose concentration such that h=1
allolactose: A Repressor: R
I h
R = h5 h5 + A5 = 1 1 + (A/h)5
repressor allolactose
Complete mathematical model
R: repressor, M: messenger & A: allolactose:
c0: basal transcription rate, c0+c: transcription rate when operon is “on”, d and δ are decay rates of mRNA and allolactose, ML is the permease mediated influx
- vMA term: B-galactosidase hydrolizes allolactose.
R=0: operon “on” R=1: operon “off”
R = 1 1 + An , dM dt = c0 + c(1 − R) − dM = c0 + cAn 1 + An − dM , dA dt = ML − δA − vMA ,
Nullclines
A M
c0 d c0+c d
M = c0 d + (c/d)An 1 + An
M = δA L − vA
A’=0: M’=0:
sigmoid Hill function Origin: M = (δ/L)A
Quasi steady state dM/dt = 0
L ¯ A
For one concentration of L three concentrations of A
Lmin Lmax
dt − − dA dt = ML − δA − vMA
M = c0 d + (c/d)An 1 + An
with
Observed bi-stability in E. coli
Green: E. coli with high expression of lac operon From: Ozbudak et al. Nature, 2004 (see the reader)
Bi-stability in growth of E. coli
The Innate Growth Bistability and Fitness Landscapes of Antibiotic-Resistant Bacteria
- J. Barrett Deris,1,2* Minsu Kim,1*† Zhongge Zhang,3 Hiroyuki Okano,1 Rutger Hermsen,1,2‡
Alexander Groisman,1 Terence Hwa1,2,3§
Science 2013
bi-stability in algal densities in lakes
a
Fold 2 Fold 1 Phosphorus input rate,
- P concentration
Oligotrophic state Eutrophic state 0.2 0.3 0.4 0.5 0.6 0.7 0.5 1 1.5 2 Bistability region
LETTER
doi:10.1038/nature11655
Flickering gives early warning signals of a critical transition to a eutrophic lake state
Rong Wang1,2, John A. Dearing1, Peter G. Langdon1, Enlou Zhang2, Xiangdong Yang2, Vasilis Dakos3,4 & Marten Scheffer3
Nature 2012
Initiation and termination of epileptic seizures
PNAS: 2012
A i C
Post-ictal critical transition
i c t a l
s e p a r a t r i x
increasing connectivity bistability
100ms 10mV
ii iii
−2
Temporal-Corr
**
Slope
10
−2
[ ]
Mid-seizure Pre-termination
MODEL
D
1.0
Proportion
0.2 0.0 Ictal Post- Pre- Ictal Post- Pre-
** ** ** **
Spatial-Corr
*
p<0.005
** * p<0.05
1.2 0.2 3400 4200
s t a b l e l . c . stable f.p. unstable l.c.
C
Post-ictal Ictal critical transition
he i c t a l
unstable f.p.
B
Frequency
Human seizures self-terminate across spatial scales via a critical transition
Mark A. Kramera,1, Wilson Truccolob,c,d,e, Uri T. Edena, Kyle Q. Lepagea, Leigh R. Hochbergc,d,e,f,g, Emad N. Eskandarf,h, Joseph R. Madseni,j, Jong W. Leek, Atul Maheshwarid,f, Eric Halgrenl, Catherine J. Chud,f, and Sydney S. Cashd,f
Initiation and termination of depression
PNAS: 2014
autocorrelation variance autocorrelation variance correlation
A C
D
E
G
F
H
I
K
J
L
B
within-valence between-valence correlation within-valence between-valence
200 8 time emotion strength 200 8 time emotion strength
x1, x2 x3, x4 x1, x2 x3, x4
x1 x2 x3 x4 0.6 SD 0.5 1.5 0.5 1.5 x1(t) / x1 x1(t+1) / x1 x1 x2 x3 x4 0.6 SD 0.5 1.5 0.5 1.5 x1(t) / x1 x1(t+1) / x1 0.5 1.5 0.5 1.5 x1(t) / x1 x2(t) / x2 0.5 1.5 0.5 1.5 x1(t) / x1 x3(t) / x3 0.5 1.5 0.5 1.5 x1(t) / x1 x2(t) / x2 0.5 1.5 0.5 1.5 x1(t) / x1 x3(t) / x3 AR(1)=0.38 AR(1)=0.77 =0.29 =-0.47 =0.69 =-0.83
Critical slowing down as early warning for the onset and termination of depression
Ingrid A. van de Leemputa,1,2, Marieke Wichersb,1, Angélique O. J. Cramerc, Denny Borsboomc, Francis Tuerlinckxd, Peter Kuppensd,e, Egbert H. van Nesa, Wolfgang Viechtbauerb, Erik J. Giltayf, Steven H. Aggeng, Catherine Deromh,i, Nele Jacobsb,j, Kenneth S. Kendlerg,k, Han L. J. van der Maasc, Michael C. Nealeg, Frenk Peetersb, Evert Thieryl, Peter Zacharm, and Marten Scheffera
2001
- Fraction of lake surface covered
by charophyte vegetation Total P (mg l –1) 0.1 0.2 0.3 0.4 0.5 0.05 0.10 0.15 0.20 0.25 0.30
Ecosystem state Conditions Perturbation F1 F2