SLIDE 1
Power, Sample Size, and the FDR
Peter Dalgaard
Department of Biostatistics University of Copenhagen
Center for Bioinformatics, Univ.Copenhagen, June 2005
Sample Size
“How many observations do we need?” Depends on
- Design
- Standard error of measurements
- Effect size
- How sure you want to be of finding it
Reminders
(Continuous data) One-sample (or paired, differences): SEM = s ×
- 1/n
Significance if |¯ x − µ0| SEM > t.975(DF) Two-sample: SEDM = s ×
- 1/n1 + 1/n2
|¯ x1 − ¯ x2| SEDM > t.975(DF) t.975(DF) ≈ 2. Notice that SE(D)M decreases with n.
Variation of Observations and Means
−3 −2 −1 1 2 3 0.0 0.5 1.0 1.5 x dnorm(x, sd = sqrt(1/20))
t Test
−3 −2 −1 1 2 3 0.0 0.1 0.2 0.3 0.4 t dt(t, 38)
- If there is no (true) difference then there is little chance of
getting an observation in the tails
- If there is a difference, then the center of the distribution is
shifted.
Type I and Type II Errors
A test of a hypothesis can go wrong in two ways: Type I error: Rejecting a true null hypothesis Type II error: Accepting a false null hypothesis Error probabilities: α resp. β α: Significance level (0.05, e.g.) 1 − β: Power – probability of detecting difference Notice that the power depends on the effect size as well as on the number of observations and significance level.
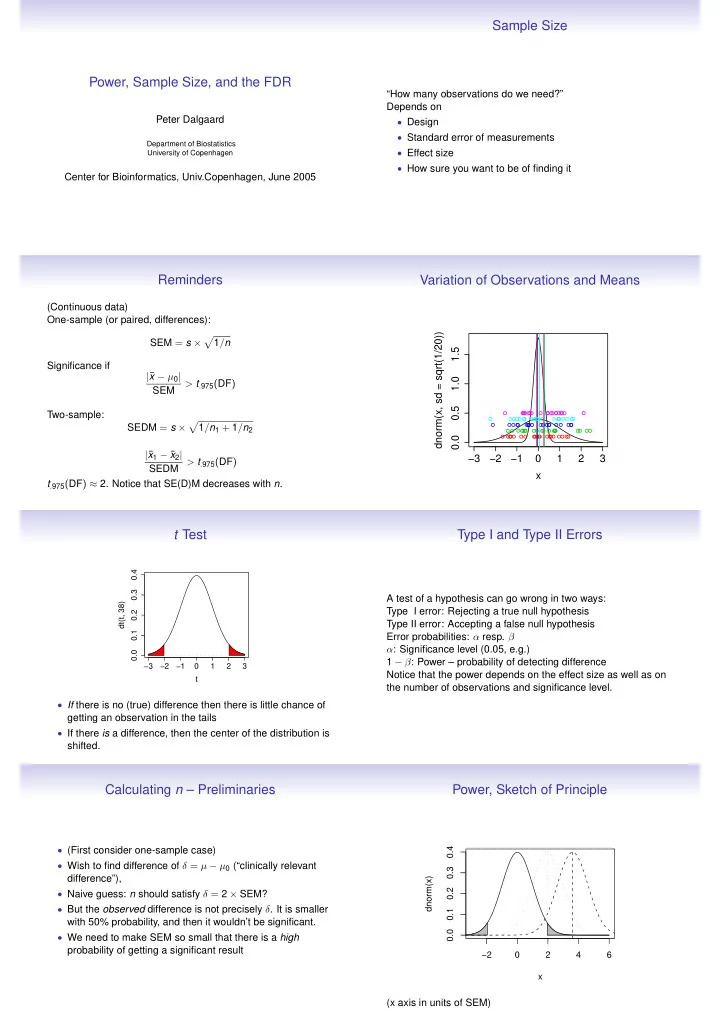
Calculating n – Preliminaries
- (First consider one-sample case)
- Wish to find difference of δ = µ − µ0 (“clinically relevant
difference”),
- Naive guess: n should satisfy δ = 2 × SEM?
- But the observed difference is not precisely δ. It is smaller
with 50% probability, and then it wouldn’t be significant.
- We need to make SEM so small that there is a high