SLIDE 1
Joint and marginal probabilities Joint: Marginal: How to compute - - PowerPoint PPT Presentation
Joint and marginal probabilities Joint: Marginal: How to compute - - PowerPoint PPT Presentation
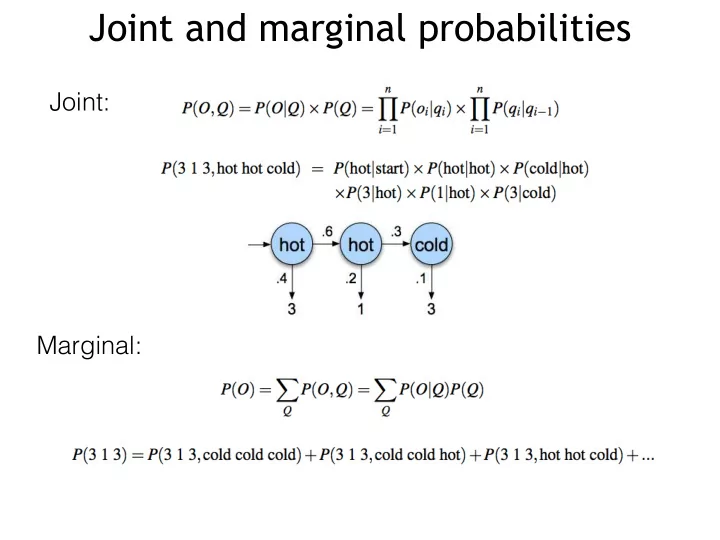
Joint and marginal probabilities Joint: Marginal: How to compute the probability of observations Forward algorithm Forward algorithm Forward algorithm Decoding: finding the most probable states Similar to the forward algorithm, we can define
SLIDE 2
SLIDE 3
Forward algorithm
SLIDE 4
Forward algorithm
SLIDE 5
Forward algorithm
SLIDE 6
Decoding: finding the most probable states
Similar to the forward algorithm, we can define the following value:
SLIDE 7
SLIDE 8
Viterbi algorithm
SLIDE 9
Gene finding
SLIDE 10
Gene finding
SLIDE 11
- In human genome, ~3% of DNA sequence is
genes
- Lot of “junk” DNA between genes, and even
inside genes (between exons).
- Due to the reverse complement, one gene can
start from either direction.
- Gene finding must deal with these.
Gene finding
SLIDE 12
In bacteria, there is no intron in the coding region.
Gene finding for bacterial genomes
SLIDE 13
Gene finding for bacterial genomes
In bacteria, there is no intron in the coding region. Codon usage can be different between the noncoding regions and coding regions.
start state stop state
SLIDE 14
Codon usages can be different in the typical coding regions and the atypical coding regions. Horizontal gene transfer
Gene finding for bacterial genomes
SLIDE 15
Gene finding: frames
SLIDE 16
Gene finding: HMM for bacteria
GeneMarker’s HMM model
SLIDE 17
Gene finding: handling introns
SLIDE 18
Gene finding: handling introns
Splicing site motifs
SLIDE 19
Splicing site motifs
Gene finding: handling introns
SLIDE 20