Toward comprehensive whole-cell models Genomic and biochemical data - - PowerPoint PPT Presentation
Toward comprehensive whole-cell models Genomic and biochemical data - - PowerPoint PPT Presentation
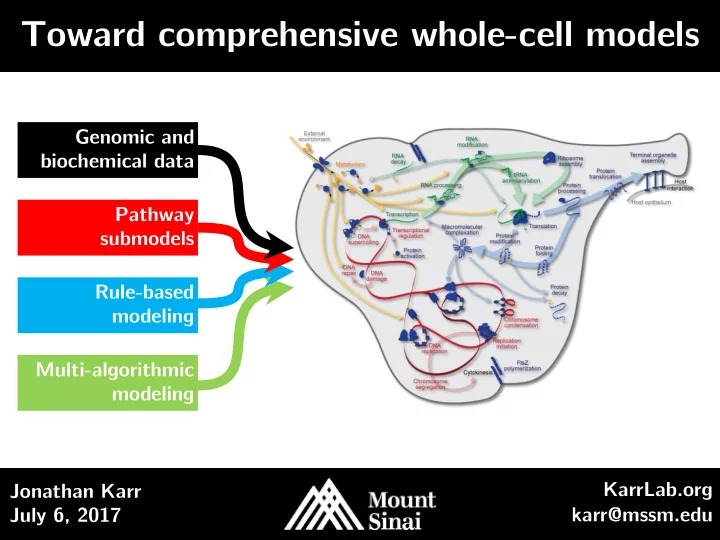
Toward comprehensive whole-cell models Genomic and biochemical data Pathway submodels Rule-based modeling Multi-algorithmic modeling KarrLab.org Jonathan Karr July 6, 2017 karr@mssm.edu Outline Introduction to whole-cell (WC) modeling
Outline
Introduction to whole-cell (WC) modeling
- What is a WC model?
- Motivation
- Challenges
- Feasibility
Methodology
- Data aggregation
- Data organization
- Hybrid simulation
New tools to accelerate WC modeling
- Data aggregation
- Model representation
- Parallel simulation
Outline
Introduction to whole-cell (WC) modeling
- What is a WC model?
- Motivation
- Challenges
- Feasibility
Methodology
- Data aggregation
- Data organization
- Hybrid simulation
New tools to accelerate WC modeling
- Data aggregation
- Model representation
- Parallel simulation
Features of whole-cell (WC) models
Karr et al., 2015
Whole organism Dynamic Whole genome including each gene Stochastic Whole cell cycle Accurate Species-specific Mechanistic
AGTC
Outline
Introduction to whole-cell (WC) modeling
- What is a WC model?
- Motivation
- Challenges
- Feasibility
Methodology
- Data aggregation
- Data organization
- Hybrid simulation
New tools to accelerate WC modeling
- Data aggregation
- Model representation
- Parallel simulation
Genome design requires WC models
Tissue engineering Biosensors Biofactories
Example: drug biosynthesis
Example: drug biosynthesis
Example: drug biosynthesis
Example: drug biosynthesis
Example: drug biosynthesis
Example: drug biosynthesis
Example: drug biosynthesis
Personalized medicine requires WC models
Outline
Introduction to whole-cell (WC) modeling
- What is a WC model?
- Motivation
- Challenges
- Feasibility
Methodology
- Data aggregation
- Data organization
- Hybrid simulation
New tools to accelerate WC modeling
- Data aggregation
- Model representation
- Parallel simulation
WC models are a grand challenge
Time Length
Replication Growth Transcription Metabolism
Challenge: multiple time and length scales
Single-cell variation Microscopy Transcription RNA-seq Protein expression Mass-spec, Western blot
Challenge: heterogeneous data
Challenge: sparse data
Metabolic Signaling Transcriptional regulatory
Challenge: heterogeneous granularity
Outline
Introduction to whole-cell (WC) modeling
- What is a WC model?
- Motivation
- Challenges
- Feasibility
Methodology
- Data aggregation
- Data organization
- Hybrid simulation
New tools to accelerate WC modeling
- Data aggregation
- Model representation
- Parallel simulation
WC modeling is now feasible
Genomic and biochemical data Pathway submodels Rule-based modeling Multi-algorithmic modeling
Extensive molecular data is available
Numerous predictors are available
- miRNA targets: TargetScan
- Operons: OperonPredictor
- Protein half-lives: N-end rule
- Protein localization: PSORT
- Signal sequences: SignalP
- Transcription start site: Promoter
Numerous databases are available
Model design tools are available
MetaFlux
Model languages are available
Numerous pathway models are available
Numerous simulators are available
Uptake FBA Composition Metabolism FBA Composition Transcription Stochastic binding Gene expression Translation Stochastic binding Gene expression Replication Chemical kinetics DNA sequence
Testing tools are available
PRISM
Numerous other tools
- Automated model construction
- Model refinement
- Parallel simulation
- Calibration
- Analysis and visualization
- …
Outline
Introduction to whole-cell (WC) modeling
- What is a WC model?
- Motivation
- Challenges
- Feasibility
Methodology
- Data aggregation
- Data organization
- Hybrid simulation
New tools to accelerate WC modeling
- Data aggregation
- Model representation
- Parallel simulation
Pathway modeling workflow
- 1. Choose a system to model
- 2. Determine the scope and granularity of the model
- 3. Determine the mathematical representation of the
model
- 4. Reconstruct the species, reactions, rate laws, and
rate parameters from the literature
- 5. Debug and calibrate the model by comparison to
data
- 6. Test the model by comparison to independent data
Detail Scope
ODE Shuler, 1970’s FBA Palsson, 1990’s Boolean Bolouri, 2000’s Gillespie Luthey-Schulten, 2011 PDE
WC model
Predictive modeling methodologies
Scaling pathway modeling to whole-cells
- Aggregate more data
– Accelerate data aggregation through automation – Organize input data using pathway/genome databases
- Build models collaboratively using web-based tools
– Define the semantic meaning of every model component – Track every assumption and data source
- Describe models clearly
– Explicitly describe the data used to build models – Describe models in terms of rules
- Describe and simulate hybrid models
Scaling pathway modeling to whole-cells
Genomics, bioinformatics ↔ Mechanistic modeling Pathway/genome databases ↔ Model design tools Polymers, sequences ↔ Rule-based modeling Stochastic modeling ↔ Steady-state modeling (FBA) Numerical simulation ↔ Big data analytics Model design tools ↔ Collaboration tools
WC modeling workflow
Aggregate data
Fraser et al., 1995; Kühner et al., 2009; Lluch-Senar et al., 2013; Maier et al., 2013; Yus et al. 2012
Proteome Mass-spectrometry Transcriptome RNA-seq Epigenome Meth-seq Genome DNA-seq Metabolome Mass-spectrometry
Karr et al., 2013
Organize input data
Free Bound Promoter Bound Active
- 1. Update RNA polymerase states
- 3. Bind RNA polymerase
- 2. Calculate promoter affinities
- 4. Elongate and terminate transcripts
AUGAUCCGUCUCUAAUGUCUAC UTCAACGUGAGGUAAUAAAGUC UCCACGAUGCUACUGUAUC GCCUCAUACUGCGGAU UUACGUAUCAGUGAUCAGUACU Sequence Transcript
HcrA Spx Fur GntR LuxR glpF dnaJ dnaK gntR trxB polC
Design submodels
Design pathway submodels
Uptake FBA Composition Metabolism FBA Composition Transcription Stochastic events Gene expression Translation Stochastic events Gene expression Replication Chemical kinetics DNA sequence
Uptake FBA Composition Metabolism FBA Composition Transcription Stochastic events Gene expression Translation Stochastic events Gene expression Replication Chemical kinetics DNA sequence
Submodels States
Mass, shape Metabolite, RNA, protein counts Mammalian host Transcript, polypeptide sequences DNA polymerization, proteins, modifications FtsZ ring
Combine submodels
1 s
Uptake Metabolism Transcription Translation Replication
Cell states Cell states
Uptake Metabolism Transcription Translation Replication
Cell states
Uptake Metabolism Transcription Translation Replication
Concurrently integrate submodels
Calibrate model
1.Estimate individual parameters 2.Generate reduced models of individual pathways and to calibrate individual pathways 3.Refine joint parameter values using full models
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
Verify model against known biology
Matches training data
Cell mass, volume Biomass composition RNA, protein expression, half-lives Superhelicity
Matches published data
Metabolite concentrations DNA-bound protein density Gene essentiality
Matches theory
Mass conservation Central dogma Cell theory Evolution
No obvious errors
Plot model predictions Manually inspect data Compare to known biology
- E. coli
- M. genitalium
Genome 4700 kb 580 kb Genes 4461 525 Size 2 μm × 0.5 μm 0.2-0.3 μm
Mycoplasmas are small in size and complexity
Karr et al., 2012
Model includes 28 pathway submodels
Karr et al., 2012
Submodels are integrated through 16 common states
Model represents 76% of genes
1 10 100
Genes Process
DNA RNA Protein Other
Condensation Segregation Damage Repair Replication Rep Init Trans Reg Degradation Modification Processing Transcription Aminoacylation Complexation Activation Degradation Folding Modification Processing I Processing II Translocation Ribosome Term Org Translation Shape Cytokinesis FtsZ Metabolism
v v
Karr et al., 2012
Predict energy consumption
WC models predict ancestral similarity
Optimal gene expression
- M. genitalium
- M. mycoides
- M. pneumoniae
Optimal architecture retains robustness
Optimal gene expression retains robustness
Purcell et al., 2013
WC models can inform synthetic designs
Kazakiewicz et al., 2015
WC models can reposition antibiotics
Outline
Introduction to whole-cell (WC) modeling
- What is a WC model?
- Motivation
- Challenges
- Feasibility
Methodology
- Data aggregation
- Data organization
- Hybrid simulation
New tools to accelerate WC modeling
- Data aggregation
- Model representation
- Parallel simulation
Current limitations
- M. genitalium model is limited and inaccurate
- Ignores several pathways
- Mispredicts the growth rates of many single gene disruptions
Methods are not rigorous
- Data selection
- Multi-algorithm simulation
- Parameter estimation
- Verification
Methods are time-consuming
- Data aggregation
- Model design
- Model verification
Hard to understand, reuse, reproduce
Technology development goals
Scale to more complex models Accelerate WC model building and simulation Enable more researchers to engage in WC modeling Apply WC modeling to bioengineering and medicine
WC modeling process
Accelerate data aggregation
- Chaperones
- Complex composition
- DNA binding sites
- DNA footprints
- DNA methylation
- DNA sequence
- Gene-drug interactions
- Genome annotation
- Growth rates
- Metabolite concentrations
- Protein cofactors
- Protein expression
- Protein half-lives
- Protein localization
- Protein modification
- RNA editing
- RNA expression
- RNA half-lives
- RNA modification
- RNA maturation
- Reaction fluxes
- Reaction kinetics
- Reaction stoichiometries
- Signaling pathways
- DNA mutations
Accelerate data aggregation
- Help modelers quickly get relevant data for a
model
- Enable modelers to aggregate data
collaboratively
- Record the provenance of all data
Yosef Roth
Accelerate data aggregation
- 1. Merge data from as many sources as
feasible
- 2. Find most relevant data for the model
- Species: taxonomic distance
- Environment: temperature, pH, media
- 3. Normalize data
- 4. Calculate weighted consensus of the
relevant data
- 5. Record all provenance
Accelerate data aggregation
Accelerate data aggregation
- Metabolite concentrations: ECMDB, YMDB
- RNA expression: Array Express
- Protein expression: PaxDB
- Protein complexes: CORUM
- Protein localization: prediction
- Protein-DNA interactions: DBD, DBTBS
- Reaction kinetics: SABIO-RK
Organize data for model building
Organize data for model building
Systemize model descriptions
John Sekar
Systemize model descriptions
Initiation
Dna(pos=sample(d.tss.pos)) + RnaPol -> Dna(pos=.).RnaPol.Rna(seq=‘’) algorithm: SSA rate law: constants: kcat value = d.tss.rate refs: [10.1093/bioinformatics/btw598, …] units = 1/s
Elongation
Dna(pos=<i>).RnaPol.Rna(seq=<j>) + RevComp(DsDna(pos=<i>)) -> DsDna(pos=<i+1>).RnaPol.Rna(seq=<j>+RevComp(DsDna(pos=<i>))) x-ref: EC: x.x.x.x
84
Rule-based modeling Genomic data Bioinformatic calculations Annotation, provenance Multi-algorithmic modeling
John Sekar
James Faeder, U Pitt
Language enables compact model descriptions
Initiation Elongation Termination SBML 1 per RNA 335 1 per base ~500k 1 per RNA 335 BioNetGen 1 per RNA 335 1 per RNA 335 1 per RNA 335 WC-Lang 1 1 1 1 1 1
85
Provenance tracking
Submodel design
WC-ML, SBML, CellML
Systemize simulation
Goldberg et al., 2016
Arthur Goldberg
High-performance simulator
Simulation results database
Visual analysis of simulation results
Future work
- Online platform for collaborative model
design
- Parallel, rule-based simulator
- Scalable methods for calibrating and
validating large models
- Community standard for verifying WC
models
Karr Lab overview
Technology development Modeling language
- Programmatic
- Rule- and sequence-based
- Multi-algorithmic
Parallel simulation
- Reusable
- Multi-algorithmic
- Parallel discrete event
simulation
Pilot models
- M. pneumoniae
- Expand scope
- Improve accuracy
- Drive genome design
Stem cells
- Personalized models
- Precision medicine
WC- Rules
Summary
Genomic and biochemical data Pathway submodels Rule-based modeling Multi-algorithmic modeling
Methods development James Faeder, U Pitt
- M. pneumoniae
Veronica Llorens, CRG Maria Lluch-Senar, CRG Samuel Miravet, CRG Luis Serrano, CRG
- B. subtilis
Pablo Meyer, IBM
Acknowledgements
John Sekar Yosef Roth Roger Rodriguez Arthur Goldberg Yin Hoon Chew Balazs Szigeti