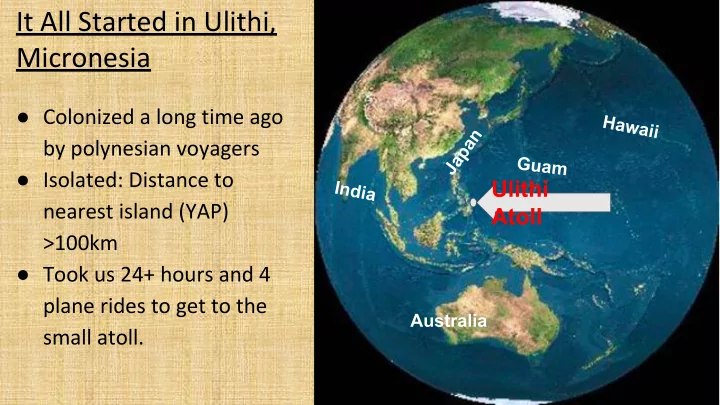
It All Started in Ulithi, Micronesia
- Colonized a long time ago
by polynesian voyagers
- Isolated: Distance to
nearest island (YAP) >100km
- Took us 24+ hours and 4
plane rides to get to the small atoll. Ulithi Atoll
Australia Japan India G u a m Hawaii
It All Started in Ulithi, Micronesia Colonized a long time ago - - PowerPoint PPT Presentation
It All Started in Ulithi, Micronesia Colonized a long time ago Hawaii Japan by polynesian voyagers G u a m Isolated: Distance to Ulithi India nearest island (YAP) Atoll >100km Took us 24+ hours and 4 plane rides to get
Australia Japan India G u a m Hawaii
Applied Research Technologies: “Genetic Barcoding of Micronesian Fish”
B Transition...
To enable screen reader support, press Ctrl+Alt+Z To learn about keyboard shortcuts, press Ctrl+slash
One People One Reef program, which is a collaboration between scientists and local communities in Micronesia for sustainable
clippings collected by local fisherman in Micronesia.
locally
Cabrillo College’s campus by students
PCR machine for Cabrillo
UC Berkeley
○ Clean genetic sequences ○ ID species (Match genetic sequences to online databases) ○ Evaluate evolutionary relationships
humans called MT-CO1)
to tell species apart
Sample:
○ Quality ○ Age ○ Made it hard to amplify the DNA ○ Tested by using gel to see quality of DNA - showed some degradation
○ Fish ○ Corals ○ Sea cucumbers
PCR troubleshooting:
for different species
temperatures for each species
Human Error:
○ Working with several samples at once
○ Microscopic amounts
○ Fish, sea cucumbers & corals
database!
amplified sea cucumber DNA ○ The COI gene of Holothuria atra ○ To be continued next semester!
Bothus mancus Flowery flounder Gymnosarda unicolor Dogtooth tuna Kyphosus vagiensis Brassy chub Parupeneus barberinus
Dash-and-dot goatfish
Siganus punctuatus Gold spotted spinefoot Stolephorus andhraensis (?) Anchovy species Siganus stellatus Brown spotted spinefoot
Thunnus albacares Yellowfin tuna Lethrinus rubrioperculatus Spotcheek Emperor Plectropomous laevis Blacksaddled coral grouper Acanthurus lineatus Lined surgeonfish
Katsuwonus pelamis Skipjack tuna Juveniles! Benthosema fibulatum Lanternfish
Cheilopogon abei Abe’s flyingfish
Identified species:
Acanthurus triostegus Convict surgeonfish Acanthurus lineatus Lined surgeonfish Caranx lugubris Black jack Caranx sexfasciatus Bigeye trevally Crenimugil crenilabis Fringelip mullet Kyphosus cinerascens Blue sea chub Lethrinus rubrioperculatus Spotcheek Emperor
Identified species (continued):
Variola albimarginata White edged lyretail Variola louti Yellow edge lyretail Thunnus albacares Yellowfin tuna Plectropomus laevis Blacksaddled coral grouper Myripristis berndti Blotch eye soldierfish Lutjanus kasmira Blue striped snapper Lutjanus gibbus Humpback red snapper Oxycheilinus unifasciatus Ring tail maori wrasse
Phylogenetic tree of a Bacteria???
Stolephorus andhraensis (?)
Potential cryptic species
Parupeneus barberinus
Dash-and-dot goatfish
identified, are the ones reported, giving their database more credibility
across all research angles (surveys, genetics, fisheries data)
found appear to be either new species or cryptic
highlight unique genetic differences of the fish in the Micronesian region
that can help guide future research
inform sustainable management
laboratory
analyze data
○ Troubleshooting ○ Team building
Cabrillo College Biology Teachers:
UCSC Professor Giacomo Bernardi
column.
proteins also
column. We now have extracted DNA!!!
homologous gene found in all species.
found in animals is high enough that species can be differentiated.
sequenced (658 base pairs, depending on the species)
Geneious
sequences
alignment; cleaned and trimmed results to create consensus sequence
Phylogenetic tree of Thunnus albacares and closely related species
to determine what species we have by matching COI sequences
and created neighbor- joining phylogenetic trees
procedure/results
expectations
techniques