Burn wounds Burns are one of the most common and devastating forms - - PowerPoint PPT Presentation
Burn wounds Burns are one of the most common and devastating forms - - PowerPoint PPT Presentation
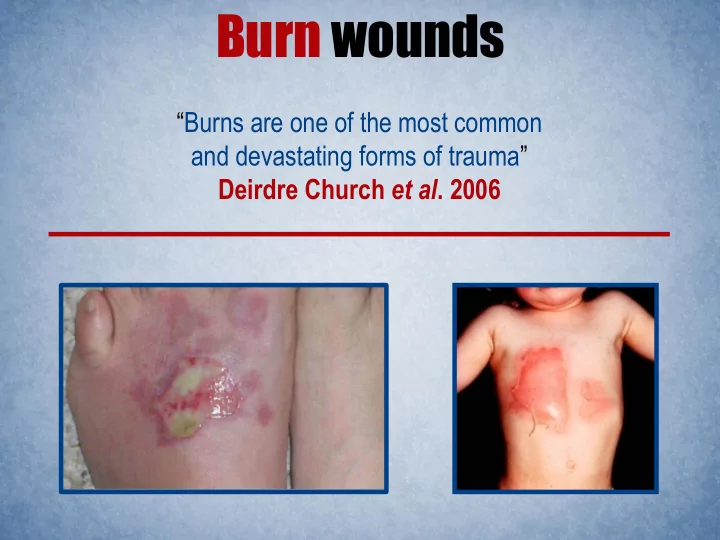
Burn wounds Burns are one of the most common and devastating forms of trauma Deirdre Church et al . 2006 Burn wounds 1,200,000 burn injuries 100,000 hospitalized 5,000 deaths 75% are infection related National Center for Injury
Burn wounds
burn injuries 1,200,000
National Center for Injury Prevention and Control in the United States
hospitalized 100,000 deaths 5,000 are infection related 75%
iGEM Groningen 2014
the smart bandage
Conventional treatments
Crèmes and bandages Bathing Preventive antibiotics Skin transplantation
Requirements
the smart bandage
- Must be used for several days
- Must detect infections
- Must secrete infection preventing molecules (IPMs)
Pseudomonas aeruginosa Staphylococcus aureus
Opportunistic pathogens
Gram-positive Gram-negative
“A post-antibiotic era - in which common infections and minor injuries can kill - far from being an apocalyptic fantasy, is instead a very real possibility for the 21st Century.” WHO, April 2014
World Health Organization (2014) Antimicrobial resistance: global report on surveillance. ISBN 978 92 4 156474 8
Our goal
To detect and prevent further infections caused by S. aureus and P. aeruginosa in burn wounds.
Why Lactococcus lactis ?
- Harmless species
- Food approved
- Produces lactate
- No spore formation
- Can be temporarily inactivated
Image adapted from a SEM scan by Joseph A. Heintz, University of Wisconsin-Madison.
Commission of Genetic Modification
Thomas and Lianne at COGEM.
How does it work?
DETECTION Quorum sensing molecules AHL (P. aeruginosa) AIP-1 (S. aureus)
Contreras, G. et al. (2013) LaSarre, B. and Michael J.F. (2013)
How does it work?
BIOFILM DESTRUCTION DspB degrades the biofilms of both species This way the other infection prevention molecules can reach them
Mark, B.L. et al. (2001)
How does it work?
QUORUM SENSING DISRUPTION AiiA inhibits AHL Quorum sensing of P. aeruginosa is disrupted
- P. aeruginosa expresses less virulence genes
Body can clear P. aeruginosa without problems
Kim, M.H. et al. (2005)
How does it work?
NISIN PRODUCTION Nisin kills Gram-positive bacteria
Wiedemann, I. et al. (2001)
Biobricks
Toolbox
- Nisin biobricks
- sfGFP
LactoAid
- AiiA
- DspB
- Gene constructs
From bacterium to bandage
Bandage materials
Top layer - polymethyl pentane
- Permeable to gases
- iGEM 2012, Groningen
Bandage materials
Middle layer – polyacrylamide gel
- Pore size is adjustable
- Nutrients can be added
- Cheap
- Rehydratable
Bandage materials
Bottom layer – cellulose nitrate
- Permeable to small molecules
and proteins
- iGEM 2013, TU Delft
Description of the model
Single unit Total model
Parameters – Rate Equations
Nisin production assay
Optimal concentration 2% Glucose
Models
Model-based design
Least optimal design (0 - 24 h) Max IPMs No IPMs
Model-based design
Best design (0 - 24 h) Max IPMs No IPMs
Modeling outcome
Time to reach effective concentrations nisin aiiA dspB 12 min 18 min 18 min 24 min 30 min 30 min 114 min 138 min 144 min
Modeling outcome
Time to reach effective concentrations nisin aiiA dspB 12 min 18 min 18 min 24 min 30 min 30 min 114 min 138 min 144 min
Modeling outcome
Time to reach effective concentrations nisin aiiA dspB 12 min 18 min 18 min 24 min 30 min 30 min 114 min 138 min 144 min
Results: Cell growth in the active layer
Results: Nisin Secretion
- Nisin-sensitive strain plated
- LactoAid active layer,
with nisin-producing strain
In conclusion
“… this is an application that appeals, it would be a waste if this idea would remain at -80°C…”
We would like to thank:
Thank YOU for your attention!
Wiedemann, I et al. . “peifi Bidig of Nisi to the Peptidogla Preursor Lipid II Coies Pore Foratio ad Ihiitio of Cell Wall Biosthesis for Potet Atiioti Ativit. The Journal of biological chemistry 276(3): 1772–79. Kim, Myung Hee et al. 2005. The Moleular “truture ad Catalti Mehais of a Quoru- Quenching N-Acyl-L-Homoserine Latoe Hdrolase. Proceedings of the National Academy of Sciences
- f the United States of America 102(49): 17606–11.
Mark, B L et al. 2001. Crstallographi Evidee for “ustrate-Assisted Catalysis in a Bacterial Beta-
- Hexosaminidase. The Journal of biological chemistry 276(13): 10330–37.
LaSarre, Breah, and Michael J Federle. . Eploitig Quoru “esig to Cofuse Baterial
- Pathoges. Microbiology and molecular biology reviews : MMBR 77(1): 73–111.
García-Contreras, Rodolfo, Toshinari Maeda, ad Thoas K Wood. . Resistae to Quoru- Quehig Copouds. Applied and environmental microbiology 79(22): 6840–46.
References
Rate equations
- M. Boonmee et al. Biochemical Engineering Journal 14 (2003) 127–135
Shimizu et al. Appl. Environ. Microbiol. 1999, 65(7):3134.
X1 = X2 = X3 = x7 = X8 = X9 = X10 = X11 = IPMs =
Rate equation
F (variable,parameters) Diffusion term
+
Rate =
Diffusion equations
Volume fraction of the gel: Density of IPMs:
mw - mass of the water mb - mass of the buffer mp - mass of the polymer vp - partial specific volume of gel in water vwb - partial specific volume of gel in buffer
Tong, Jane, and John L. Anderson. Biophysical journal 70.3 (1996): 1505-1513. Fischer Hannes, et al. Protein Science 13.10 (2004): 2825-2828.
Diffusion equations
Stokes-Einstein equation: Diffusion rates of the IPMs were found with:
Tong, Jane, and John L. Anderson. Biophysical journal 70.3 (1996): 1505-1513.
Model-based design
Expected most optimal design Max IPMs No IPMs
Growth rate (0.653 hr-1) 39 minutes
0.01 0.1 1 10 2 3 4 6 7 8 9 10 11
Log OD600nm Time (Hours)
Parameters – Rate Equations
Growth rate – L. lactis
Rate equation
F (variable,parameters) Diffusion term
+
Rate
=
Results: Protein production in the gel
30 min 90 min 60 min In the Gel On the Gel
Toolbox
Nisin gene cluster