1
In paramagnetic systems: µS=658.2 µI Dipole-dipole interaction:
( )( )
- ⋅
− ⋅ ⋅ − =
3 5 dip
3 4 r r E
2 1 2 1
µ µ µ µ µ µ µ µ µ µ µ µ µ µ µ µ r r π µ
R1,2
1,2 ∝
∝ ∝ ∝ ∝ ∝ ∝ ∝ <E <E2>f(τ τ τ τ τ τ τ τc
c,
, ω ω ω ω ω ω ω ω)
µS µI1 µI2
Relaxation rates:
Paramagnetic metalloproteins Paramagnetic metalloproteins
Econt ∝ AcSz
( )
- +
+ + +
- =
2 2 2 2 6 2 2 2 2 1
1 3 1 7 1 4 15 2
c I c c S c B e I M
r S S g R τ ω τ τ ω τ µ γ π µ ( )
- +
+ + + +
- =
2 2 2 2 6 2 2 2 2 2
1 3 1 13 4 1 4 15 1
c I c c S c c B e I M
r S S g R τ ω τ τ ω τ τ µ γ π µ
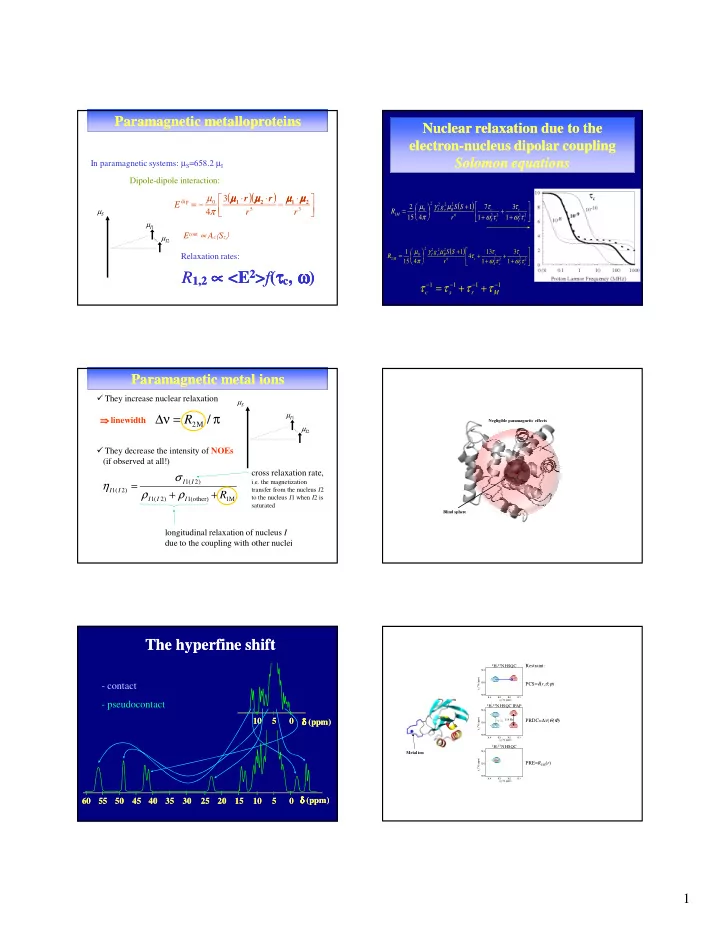
Nuclear relaxation due to the Nuclear relaxation due to the electron electron-nucleus dipolar coupling nucleus dipolar coupling Solomon equations Solomon equations
τc
1 1 1 1 − − − −
+ + =
M r s c
τ τ τ τ
µS µI1 µI2
They increase nuclear relaxation They decrease the intensity of NOEs (if observed at all!)
π = ∆ν /
M 2
R
M 1 (other) 1 ) 2 ( 1 ) 2 ( 1 ) 2 ( 1
R
I I I I I I I
+ + = ρ ρ σ η
longitudinal relaxation of nucleus I due to the coupling with other nuclei
Paramagnetic metal ions Paramagnetic metal ions
- linewidth
cross relaxation rate,
i.e. the magnetization transfer from the nucleus I2 to the nucleus I1 when I2 is saturated
Negligible paramagnetic effects Blind sphere
δ δ δ δ δ δ δ δ (ppm) (ppm) 5 10 10
The hyperfine shift The hyperfine shift
δ δ δ δ δ δ δ δ (ppm) (ppm) 5 10 10 20 20 25 25 15 15 35 35 40 40 30 30 50 50 55 55 45 45 60 60
- contact
- pseudocontact
8.4 8.3 8.2 8.1 128 126 124 δ1(
15N) (ppm)δ2(
1H) (ppm)8.4 8.3 8.2 8.1 128 126 124 δ1(
15N) (ppm)δ2(
1H) (ppm)8.4 8.3 8.2 8.1 128 126 124 δ1(
15N) (ppm)δ2(
1H) (ppm)Metal ion
1H-15N HSQC 1H-15N HSQC IPAP 94 Hz 114 Hz 1H-15N HSQC
Restraint: PCS=δ(r,ϑ,ϕ) PRDC=∆ν(Θ,Φ) PRE=R1M(r)