Vi t l N Virtual Neurons
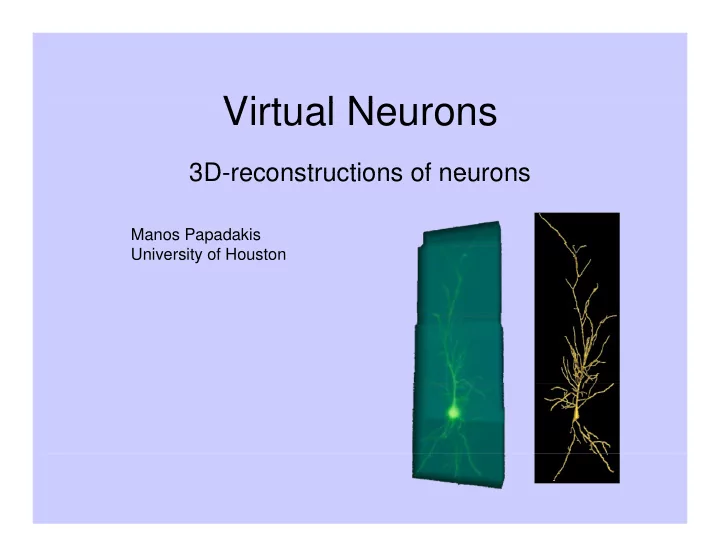
3D reconstructions of neurons 3D-reconstructions of neurons
Manos Papadakis p University of Houston
Vi t Virtual Neurons l N 3D reconstructions of neurons - - PowerPoint PPT Presentation
Vi t Virtual Neurons l N 3D reconstructions of neurons 3D-reconstructions of neurons Manos Papadakis p University of Houston Collaborators I. Kakadiaris, I. Kakadiaris, D. Labate Neuroscience/data acquisition collaborators: q P. Saggau
Manos Papadakis p University of Houston
Neuroscience/data acquisition collaborators: q
Signal In Signal Out Structure and Function
3
Do the last task when a neuron is activated and responds. This will help add a layer of a true This will help add a layer of a true activation/response model.
4
Raw Data Reconstruction
5
30 m
5 m 10 m i i t it j ti
6
x –y maximum intensity projection y –z maximum intensity projection
2 m Tubular-like 2 2 m 2 m
7
8
Input: Raw data Step 3 Dendrite Detection Step 2. Denoising Step 1. Deconvolution Step 3. Dendrite Detection
Shape learning and shape prediction 3D Frame-Based Denoising
p g
Experimental PSF
Step eco
Shape Model Shape Model
Multiple image stacks registration
Step 4. Registration
Medial axis and
Step 5. Morphological Reconstruction Step 6. Statistical analysis Output: Simulation of Computational Model
stacks registration radius estimation Hoc file generation
9
σ
5 m
10
Shape p Learning
Model to Learn Data to predict Prediction
11
Original Data Orion 1 Original Data Orion 1
12
Frangi Sato
Medium Poor
13
14
O i 1 R t ti f Di d D t S t 1 Orion 1 Reconstruction for Diadem Data Set 1
Volume 1: CF1 Volume 2: CF2
Volume 1: OP1 Volume 3: OP3
Raw data Prediction Raw data Prediction
influenced by the rotations of f ?
S b l ? e.g. Sobolev norms?
800 x 800 400 x 400
Cylinder along the x-axis for sanity check