1
Emad Tajkhorshid
Theoretical and Computational Biophysics Group Beckman I nst it ut e University of I llinois at Urbana- Champaign
Computational Chemistry GRI D Conf erence, 2003
Molecular Mechanisms of Photoactivation and Spectral Tuning in Retinal Proteins
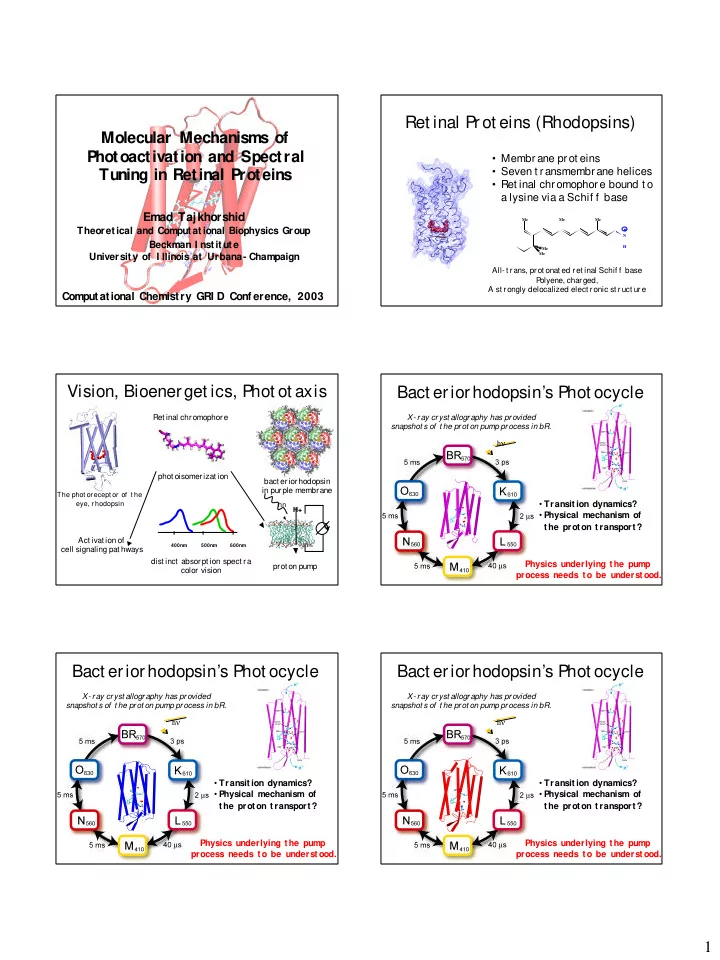
Ret inal Prot eins (Rhodopsins)
All- t rans, prot onat ed ret inal Schif f base P
- lyene, charged,
A st rongly delocalized elect ronic st ruct ure
- Membrane prot eins
- Seven t ransmembrane helices
- Ret inal chromophore bound t o
a lysine via a Schif f base
N Me Me Me Me Me H
Vision, Bioenerget ics, Phot ot axis
V H+ hn
bact eriorhodopsin in purple membrane
500nm 600nm 400nm
dist inct absorpt ion spect ra color vision
The phot or ecept or of t he eye, r hodopsin
phot oisomerizat ion Ret inal chromophore prot on pump Act ivat ion of cell signaling pat hways X- ray cryst allography has provided snapshot s of t he prot on pump process in bR.
- Transit ion dynamics?
- Physical mechanism of
t he prot on t ransport ?
hν
Physics underlying t he pump process needs t o be underst ood.
Bact eriorhodopsin’s Phot ocycle
X- ray cryst allography has provided snapshot s of t he prot on pump process in bR.
- Transit ion dynamics?
- Physical mechanism of
t he prot on t ransport ?
hν
Physics underlying t he pump process needs t o be underst ood.
Bact eriorhodopsin’s Phot ocycle
X- ray cryst allography has provided snapshot s of t he prot on pump process in bR.
- Transit ion dynamics?
- Physical mechanism of
t he prot on t ransport ?
hν
Physics underlying t he pump process needs t o be underst ood.