Microbial Genomics Microbial Genomics
Michael J. Stanhope, Michael J. Stanhope,
- Pop. Med. Diagnostic
- Pop. Med. Diagnostic Sci
Sci. .
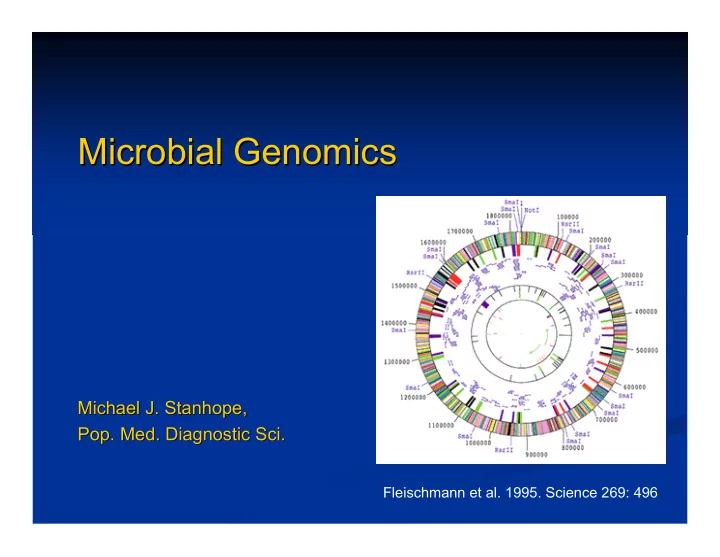
Fleischmann et al. 1995. Science 269: 496
Microbial Genomics Microbial Genomics Michael J. Stanhope, Michael - - PowerPoint PPT Presentation
Microbial Genomics Microbial Genomics Michael J. Stanhope, Michael J. Stanhope, Pop. Med. Diagnostic Sci Pop. Med. Diagnostic Sci. . Fleischmann et al. 1995. Science 269: 496 Outline Outline Introduction Introduction
Michael J. Stanhope, Michael J. Stanhope,
Sci. .
Fleischmann et al. 1995. Science 269: 496
Microbial diversity
Universal Tree of Life
Core and pan genomes
Mechanisms of HGT
Detecting HGT
http://www.ucmp.berkeley.edu/archaea/archaeamm.html
Fermentation or respiration; respire aerobically or anaerobically anaerobically; glucose or lactose as sole carbon source ; glucose or lactose as sole carbon source – – transforming sugar into amino acids, vitamins, transforming sugar into amino acids, vitamins, nucleotides nucleotides
Alcohol fermentation
Lactic acid fermentation present in eukaryotes
present in eukaryotes & prokaryotes & prokaryotes
Aerobic respiration
Oxygenic photosynthesis
Anaerobic degradation of carbohydrates through the Embden Embden-
Meyerhof pathway.
Other fermentation pathways e.g. phosphoketolase phosphoketolase pathway pathway
Anaerobic respiration
Lithotrophy ( (inorganics inorganics as source of energy) as source of energy)
Anoxygenic photosynthesis photosynthesis
Methanogenesis (H (H2
2 as energy source and produces methane)
as energy source and produces methane)
Light driven nonphotosynthetic nonphotosynthetic photophosphorylation photophosphorylation
30
Definition of species?
Lack diagnostic morphological characteristics
Exchange genetic material in unique and unusual ways
Same species = 70% DNA-
DNA hybridization
Underestimating prokaryotic diversity
Practical limitations in counting
1% cultivable
Pace, NR. 1997. Science 276:734
http://whyfiles.org/022critters/archaea.html
bacteriocytes
Distel and Cavanaugh. 1994. J. Bact. 4:1932.
Nakabachi et al. 2006. Science 314:267
(Studies on the chemical nature of the substance inducing transformations of transformations of pneumococcal pneumococcal types. J. Exp. Med.79:137)
http://fig.cox.miami.edu/Faculty/Dana/bacfun.jpg
Fixation of advantageous mutations, driven by NS =>evolutionary innovations =>evolutionary innovations
Alignable core genome size for interspecific analysis = 260
0.92 0.92 9 9 983 983 (SSI (SSI-
1, MGAS315) 0.41 0.41 4 4 978 978 (MGAS5005, M1 GAS) (MGAS5005, M1 GAS) 0.22 0.22 2 2 925 925 (MGAS9429, MGAS2096) (MGAS9429, MGAS2096) 0.00 0.00 1297 1297 SSI SSI-
1 0.00 0.00 1297 1297 M1 GAS M1 GAS 0.15 0.15 2 2 1297 1297 MGAS9429 MGAS9429 0.31 0.31 4 4 1297 1297 MGAS8232 MGAS8232 0.15 0.15 2 2 1297 1297 MGAS6180 MGAS6180 0.08 0.08 1 1 1297 1297 MGAS5005 MGAS5005 0.00 0.00 1297 1297 MGAS315 MGAS315 0.08 0.08 1 1 1297 1297 MGAS2096 MGAS2096 0.08 0.08 1 1 1297 1297 MGAS10750 MGAS10750 0.23 0.23 3 3 1297 1297 MGAS10394 MGAS10394 0.54 0.54 7 7 1297 1297 MGAS10270 MGAS10270 % under PS % under PS nbr nbr under PS under PS nbr nbr analyzed analyzed Lineage Lineage
223 (18%) 223 (18%) 18 18 (1%) (1%) 7 (1%) 7 (1%) 34 (3%) 34 (3%) 222 (18%) 222 (18%)
S.
agalactiae
477 (37%) 477 (37%) 186 186 (14%) (14%) 168 (13%) 168 (13%) 284 284 (22%) (22%) 434 (33%) 434 (33%)
S.
pyogenes
53 (20%) 53 (20%) 11 11 (4%) (4%) 35 (14%) 35 (14%) 54 54 (21%) (21%) 26 (10%) 26 (10%)
interspecific interspecific
SPI U SPI U intragenic intragenic set set SPI SPI ∩ ∩ PHI PHI PHI PHI ∩ ∩ MaxChi MaxChi ∩ ∩ NSS NSS
(set of (set of intragenic intragenic methods) methods)
PHI PHI
( (intragenic intragenic method) method)
SPI SPI (strong
(strong phylogenetic phylogenetic incongruence) incongruence)
Data set Data set
17 (53%) 17 (53%) 21 (65%) 21 (65%) 25 (78%) 25 (78%) 32 32
S.
pyogenes
4 (40%) 4 (40%) 4 (40%) 4 (40%) 10 10
S.
agalactiae
29 (11%) 29 (11%) 20 (8%) 20 (8%) 43 (25%) 43 (25%) 175 175
interspecific interspecific
PS + PS + intragenic intragenic PS + SPI PS + SPI PS + PS + recombinant recombinant Genes Genes under PS under PS Data set Data set
Significant affect of lineage (ANOVA; p<0.0001):
Majority of pairwise pairwise multiple comparisons significantly different multiple comparisons significantly different
Significant affect of biochemical category (p<0.0001)
Amino acid biosynthesis; Biosynthesis of cofactors, prosthetic groups, roups, and carriers; Cell envelope; Cellular processes; Central interme and carriers; Cell envelope; Cellular processes; Central intermediary diary metabolism; metabolism; DNA metabolism
; Energy metabolism; Fatty acid and and phospholipid phospholipid metabolism; Hypothetical proteins; Protein fate; metabolism; Hypothetical proteins; Protein fate; Protein synthesis; Protein synthesis; Purines Purines, , pyrimidines pyrimidines, nucleosides, and nucleotides; , nucleosides, and nucleotides; Regulatory functions; Signal transduction; Regulatory functions; Signal transduction; Transcription
; Transport and binding proteins; Unknown function Transport and binding proteins; Unknown function
Significant interaction between lineage and biochemical category (p=0.003) (p=0.003)
(S.
pneumoniae, S. , S. suis suis) DNA metabolism, Transcription, ) DNA metabolism, Transcription, Protein fate Protein fate
19 unique loci for S. suis; 15 for S. thermophilus; 14 for S. pneumoniae
Haemophilus influenzae 1.8 Mb Library of plasmid clones, 1600-2000 bp fragments; sequences of these clones with their many overlaps represent the raw data entered into computer programs (e.g. TIGR assembler) which assemble the genome; remaining gaps closed with other strategies (e.g. long range PCR)
Fleischmann et al. 1995. Science 269: 496
X Prize Foundation: $5 million
1000 NA Single-molecule array VisiGen Biotechnologies 500 35 Parallel microchip Solexa 100 30 Map and survey microarray NimbleGen Systems 5 800+ Biochip Network Biosystems 7 850- 1000 Parallel bead array Microchip Biotechnologies 14,000 20,000 Electronic microchip LI-COR Biosciences 3-4 1000 Capillary electrophoresis Applied Biosystems 200 50 Sequencing by ligation Agencourt Bioscience 96 100 Parallel bead array 454 Life Sciences Expected Throughput Mb (million bases)/day Read Length (bases) Format Company Searching for Cheaper Genome Sequencers
from: Service, RF 2006. Science 311:1544
from: Margulies et al. 2005 Nature 437:376
(tracks bases as they are added); pyrosequencing
1.6 million wells;
from: Margulies et al. 2005 Nature 437:376
pyrophophate, prompting luciferase & flash of light
from each cell with nucleotides presented in flow through, computer tracks sequence growth
from: Service, RF 2006. Science 311:1544
=> compare genomes, find potential drug targets shared by clinically important range of clinically important range of taxa taxa, & absent or divergent from , & absent or divergent from human host human host
http://eburst.mlst.net/6.asp
E.g. Combimatrix Combimatrix 4 X 2K 4 X 2K microarrays microarrays
http://chunlab.snu.ac.kr/meta.htm