Chapter 8: Modeling chains of ODEs d R d t = [ r (1 R/K ) bN ] R , - - PowerPoint PPT Presentation
Chapter 8: Modeling chains of ODEs d R d t = [ r (1 R/K ) bN ] R , - - PowerPoint PPT Presentation
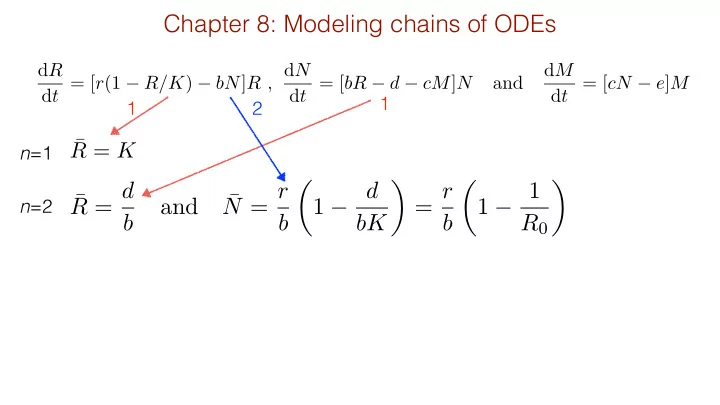
Chapter 8: Modeling chains of ODEs d R d t = [ r (1 R/K ) bN ] R , d N d M d t = [ bR d cM ] N and d t = [ cN e ] M , 1 1 2 R = K n =1 R = d N = r 1 d = r 1 1 and n =2 b b bK b R 0
Modeling chains of ODEs
dR dt = [r(1 − R/K) − bN]R , dN dt = [bR − d − cM]N and dM dt = [cN − e]M ,
¯ R = d b and ¯ N = r b ✓ 1 − d bK ◆ = r b ✓ 1 − 1 R0 ◆
¯ R = K
¯ N = e c , ¯ R = K ✓ 1 − be cr ◆ and ¯ M = b ¯ R − d c
n=1 n=2 n=3 For odd chain lengths R depends on K 1 2 3
R0
0 = cr
be
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>Modeling chains of ODEs
dR dt = [r(1 − R/K) − bN]R , dN dt = [bR − d − cM]N and dM dt = [cN − e]M , K
(a)
¯ R
KNKM
d b
n=1 n=3 n=2
(b)
¯ N
KNKM
e c
n=2 n=3
(c)
¯ M
KM
n=3
Kaunzinger & Morin, Nature, 1998
Prey alone Prey with predator Predator Bacterial food chain
Modeling chains with saturated interaction terms
dR dt = r ⇣ 1 R K ⌘
- bN
hR + R
- R ,
dN dt = bR hR + R d cM hN + N
- N , and
dM dt = cN hN + N e
- M ,
fR: R and N fN: in absence of M no N fM: no M
dR dt = r ⇣ 1 R K ⌘
- bN
hR + R + N
- R ,
dN dt = bR hR + R + N d cM hN + N + M
- N , and
dM dt = cN hN + N + M e
- M
Per capita function always depends on variable itself.
aXY ' aXY 1 + X/k + Y/k when k is large
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>Modeling chains with Beddington interaction terms
(a)
¯ R
KNKM
d b
n=1 n=3 n=2
(b)
¯ N
KNKM
e c
n=2 n=3
(c)
¯ M
KM
n=3
(d)
K ¯ R
KN KM
n=1 n=2
n=3
(e)
K ¯ N
KN KM
n=2 n=3
(f)
K ¯ M
KM
n=3
Other famous chains don’t suffer from this problem
dS dt = s − dS − βSI , dE dt = βSI − (d + γ)E , dI dt = γE − (δ + r)I and dR dt = rI − dR
SEIR model:
¯ R = r d ¯ I , ¯ I = γ δ + r ¯ E , ¯ S = (d + γ)(δ + r) γβ , ¯ E = s d + γ − d(δ + r) γβ
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>R and I are proportional to previous level _ _
dN0 dt = s − (p + d)N0 , dNi dt = 2pNi−1 − (p + d)Ni and dNn dt = 2pNn−1 − dNn ,
¯ N0 = s p + d , ¯ Ni = 2p p + d ¯ Ni−1 and ¯ Nn = 2p d ¯ Nn−1
J = −(p + d) . . . . . . 2p −(p + d) . . . . . . 2p −(p + d) . . . . . . . . . . . . . . . 2p −d
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>(J00 − λ)(J11 − λ)(J22 − λ) . . . (Jnn − λ) = 0
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>Cascade of cell divisions
Cascade of cell divisions
dN0 dt = s − (p + d)N0 , dNi dt = 2pNi−1 − (p + d)Ni and dNn dt = 2pNn−1 − dNn ,
¯ N0 = s p + d , ¯ Ni = 2p p + d ¯ Ni−1 and ¯ Nn = 2p d ¯ Nn−1
¯ N0 = s p + d , ¯ Ni = 2ipis (p + d)i+1 and ¯ Nn = s d ✓ 2p p + d ◆n
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>dQ dt = −aQ − dQQ + d X fiNi and s = aQ
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>Kinetic proofreading
Kinetic proofreading:
¯ Cn = RL Km + L ✓ k2 k−1 + k2 ◆n
dC0 dt = k1FL − (k−1 + k2)C0 , dCi dt = k2Ci−1 − (k−1 + k2)Ci and dCn dt = k2Cn−1 − k−1Cn (8.15)
F + L
k1
⌦
k−1 C
- r
dC dt = k1FL − k−1C with F = R − C gives C = RL Km + L
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>Michaelis Menten:
F + L
k1
⌦
k−1 C0 ,
Ci−1
k2
→ Ci and Ci
k−1
→ F
<latexit sha1_base64="(null)">(null)</latexit> <latexit sha1_base64="(null)">(null)</latexit> <latexit sha1_base64="(null)">(null)</latexit> <latexit sha1_base64="(null)">(null)</latexit>F = R −
n
X
i
Ci
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>with gives
Kinetic proofreading
log[k−1] log[L]
n=0 n=10 n=100 k∗
−1
High affinity Low affinity
¯ Cn = RL Km + L ✓ k2 k−1 + k2 ◆n
Cn>θ
F + L
k1
⌦
k−1 C0 ,
Ci−1
k2
→ Ci and Ci
k−1
→ F ,
F = R − C ,
where
dC0 dt = k1FL − (k−1 + k2)C0 , dCi dt = k2Ci−1 − (k−1 + k2)Ci and dCn dt = k2Cn−1 − k−1Cn (8.15)