? 190 ATP or ATP-analogues 120 + Hp DnaB ssDNA Laboratory of - PowerPoint PPT Presentation
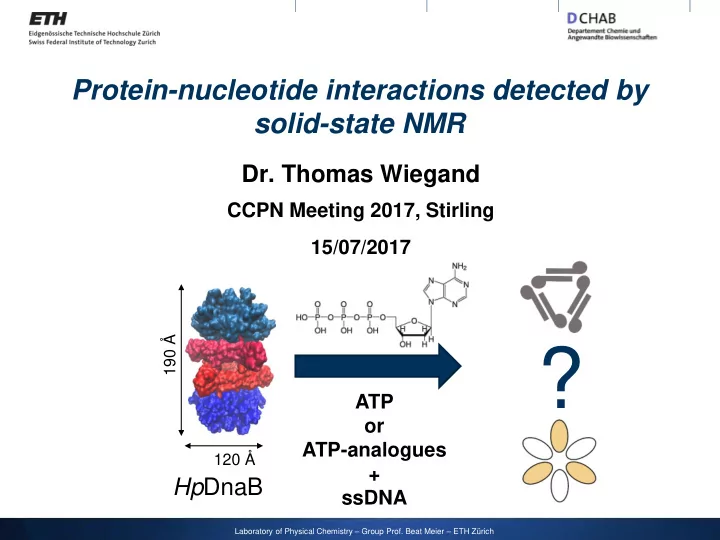
Protein-nucleotide interactions detected by solid-state NMR Dr. Thomas Wiegand CCPN Meeting 2017, Stirling 15/07/2017 ? 190 ATP or ATP-analogues 120 + Hp DnaB ssDNA Laboratory of Physical Chemistry Group Prof. Beat Meier
Protein-nucleotide interactions detected by solid-state NMR Dr. Thomas Wiegand CCPN Meeting 2017, Stirling 15/07/2017 ? 190 Å ATP or ATP-analogues 120 Å + Hp DnaB ssDNA Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich
2 DnaB helicases unwind double-stranded DNA Topoisomerase DnaG 3’ primase 5’ Parental strand DnaB helicase Taken from: Adapted from: http://love-life- http://biochem.pepperdine.edu/dokuwiki/doku.php?id=chem331:dnab_helicase science.blogspot.ch/2014/09/unzipping -of-dna.html o Structural consequences of nucleotide binding (ATP & DNA)? o Structure-function relationships in protein engines? o How does DNA replication work on a molecular level ? Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich
3 ATP-hydrolysis is coupled to molecular functioning Walker, ..., Gay, The EMBO Journal, 1982 , 1 , 945-951; Spies, DNA Helicases and DNA Motor Proteins, Springer New York, 2012 . DNA translocation
4 The helicase from Helicobacter pylori forms dodecameric assemblies SF4 helicase 6 x CTD 6 x NTD 190 Å 6 x NTD 6 x CTD Homology model for the 120 Å Hp DnaB:ADP complex (based on the Aa DnaB:ADP crystal structure) Molecular mass of 672 kDa, 488 aa / monomer Strycharska, ..., Berger, Molecular Cell , 2013 , 52 , 844-854, Bazin, …, Terradot, Nucleic Acids Res., 2015 , 43 , 8564-8576.
5 Questions to be addressed by solid-state NMR Solid-state NMR on No single-crystals available sedimented samples Do the nucleotides bind to the protein? Where do they bind? What are the structural consequences of nucleotide binding?
6 Approaches to probe protein-nucleotide interactions Diamagnetic NMR ( e.g. 31 P MAS; 13 C/ 15 N CSPs; Dynamic nuclear 15 N, 13 C NCX; 13 C, 31 P and polarization (DNP) 15 N, 31 P correlations) DNA binding to Hp DnaB ATP/DNA binding Paramagnetic NMR EPR ( e.g. substitution of the Mg 2+ cofactor ( e.g. Mn 2+ -Mn 2+ DEER) by Mn 2+ or Co 2+ )
7 Part 1 Diamagnetic solid-state NMR How to monitor nucleotide binding? How to distinguish between bound und unbound nucleotides?
8 31 P NMR to distinguish between bound and unbound nucleotides 5mM AMP-PNP + 5mM MgCl 2 31 P solution-state experiments allow to assign the resonances of AMP- PNP/AMP-PN Hp DnaB + 5mM AMP-PNP 31 P direct pulsed experiments detect + 5mM MgCl 2 AMP-PNP and AMP-PN in the solution phase of the NMR rotor Hp DnaB 31 P, 1 H cross-polarization (CP) + 5mM AMP-PNP + 5mM MgCl 2 experiments detect bound AMP-PNP in slightly different conformations Solution-phase spectra @ 7.05 T Solid-phase spectra @ 11.74 T Wiegand,...,Terradot, Böckmann, Meier, Angew. Chem. Int. Ed., 2016 , 55 , 14164-14168. Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich 8
9 31 P NMR: Does ssDNA bind to the helicase? 5mM MgCl 2 + 0.5mM ssDNA 31 P solution-state experiments allow to detect unbound ssDNA Hp DnaB 31 P direct pulsed experiments detect + 5mM AMP-PNP hydrolyzed AMP-PNP (AMP-PN) in the + 5mM MgCl 2 + 0.5mM ssDNA solution-phase of the NMR rotor Hp DnaB + 5mM AMP-PNP + 5mM MgCl 2 31 P, 1 H cross-polarization (CP) + 0.5mM ssDNA experiments detect bound ssDNA to the helicase Solution-phase spectra @ 7.05 T Solid-phase spectra @ 11.74 T Wiegand,...,Terradot, Böckmann, Meier, Angew. Chem. Int. Ed., 2016 , 55 , 14164-14168. Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich 9
10 31 P chemical shifts are very sensitive to the choice of the ATP-analogue pre-hydrolytic 31 P, 1 H cross-polarization o experiments allow to detect the “transition state” bound ATP-analogues. o Structural inhomogeneities observed upon AMP-PNP binding. post-hydrolytic o ATP gets hydrolyzed during rotor filling. 31 P, 1 H CP-MAS NMR @ 11.74 T
11 DNA-binding monitored by 31 P, 1 H cross-polarization experiments DNA binding ) ( pre-hydrolytic “transition state” DnaB + ATP-analogue + ssDNA post-hydrolytic 31 P, 1 H CP-MAS NMR @ 11.74 T
12 Arginine sidechains of the protein bind to ssDNA 15 N- 13 C NCX @ 850 MHz phosphate backbone of DNA arginine sidechain
13 CSPs indicate nucleotide binding & conformational switch of the CTD + 3D NMR experiments Hp DnaB Hp DnaB + AMP-PNP + MgCl 2 Chemical-shift perturbations (CSPs) o Consequence of nucleotide-binding o Related to allosteric effects 13 C- 13 C 20 ms DARR @ 850 MHz CSP > 0.3 ppm Wiegand,...,Terradot, Böckmann, Meier, Angew. Chem. Int. Ed., 2016 , 55 , 14164-14168.
14 How to assign such a large system? Building block approach δ 1 ( 13 C)/ ppm 13 C- 13 C 20 ms DARR @ 850 MHz NTD 1-153 + ? CTD 152-488 = FL Are the domains conserved? Transfer of assignments between NTD (assignment δ 2 ( 13 C)/ ppm available) and FL Hp DnaB Wiegand, Gardiennet, .., Terradot, Böckmann, Meier, Biomol. NMR Assign. , 2016 , 10 , 13-23; possible? Tuesday, 08/09/15 Wiegand, Gardiennet, ...,Terradot, Böckmann, Meier, J. Biomol. NMR, 2016 , 65 , 79-86. Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich 14
15 Building block approach: Assignment of the full-length protein Averaged absolute 13 C and 15 N chemical shift differences between NTD and FL Hp DnaB Structural differences between isolated NTD and NTD in FL DnaB And the CTD ? Sequential assignment of the full-length protein Wiegand, Gardiennet, ...,Terradot, Böckmann, Meier, J. Biomol. NMR, 2016 , 65 , 79-86. Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich 15
16 Part 2 Paramagnetic solid-state NMR to determine protein-ATP interactions Where does the metal ion bind?
17 Paramagnetic solid-state NMR allows to identify residues in NBD Mn 2+ : Paramagnetic relaxation enhancements (PREs) Co 2+ : Pseudo-contact shifts (PCSs) I para /I dia ratio for 2D 13 C- 13 C o DARR B 0 =20.0 T o T 1e (Mn 2+ )= 30 ns o T 1e (Co 2+ )= 0.1 ns o I CP , I t1 , I t2 , I DARR o experimental conditions o Tamaki, ..., Demura, J. Biomol. NMR 2016 , 64 , 87-101.
18 NMR: Diamagnetic Mg 2+ can be substituted by paramagnetic Mn 2+ ions 3 types of resonances : 13 C- 13 C 20 ms DARR @ 850 MHz a) Not-influenced by AMP-PNP:Mn 2+ binding ( e.g. 24A) b) Attenuated upon AMP-PNP:Mn 2+ binding ( e.g. 228A) c) Broadened beyond detection upon AMP-PNP:Mn 2+ binding ( e.g. 203A, 351A) Wiegand, Lacabanne, Keller...,Terradot, Jeschke, Böckmann, Meier, Angew. Chem. Int. Ed., 2017 , 56 , 3369-3373. Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich 18
19 How to determine PREs with CCPN? o Assign 3D spectra of diamagnetic protein sample o Scale the spectra (use a resonance not affected by PREs, here 34V ) o Assign 3D spectra of paramagnetic protein sample o Extract intensities from peak assignment lists Wiegand, Lacabanne, Keller...,Terradot, Jeschke, Böckmann, Meier, Angew. Chem. Int. Ed., 2017 , 56 , 3369-3373. Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich 19
20 Site-specific determination of PREs from 3D experiments Residues with effective distances < 15 Å (distance between C α and the two nearest metal centers) Wiegand, Lacabanne, Keller...,Terradot, Jeschke, Böckmann, Meier, Angew. Chem. Int. Ed., 2017 , 56 , 3369-3373. Laboratory of Physical Chemistry – Group Prof. Beat Meier – ETH Zürich 20
21 Paramagnetic solid-state NMR allows to identify residues in NBD Long-range distance information (> 20 Å) becomes accessible, nucleotide binding domains are identified. Wiegand, Lacabanne, Keller...,Terradot, Jeschke, Böckmann, Meier, Angew. Chem. Int. Ed., 2017 , 56 , 3369-3373.
22 Part 3 31 P, 13 C correlation experiments to probe protein-nucleotide interactions Where do the ATP-analogues and DNA bind?
23 MAS-DNP for studying protein-DNA interactions 31 P- 13 C/ 15 N correlations suffer from weak NMR signal Taken from: http://www.coperetgroup.ethz.ch/research/dynam Dynamic nuclear polarization ic-nuclear-polarization--dnp--.html (DNP)- enhanced MAS Collaboration with Prof. C. Copéret (ETH Zürich)
24 MAS-DNP for studying protein-DNA interactions Hp DnaB:ADP:ssDNA ρ = (SN DNP /SN NMR ) Highest sensitivity without d8-glycerol. 2 mM AMUpol radical concentration (not optimized). Ravera, ..., Griffin, Luchinat, Bertini, J. Am. Chem. Soc., 2013 , 135 , 1641-1644; Wiegand, ..., Copéret, Böckmann, Meier, J. Biomol. NMR , submitted. Collaboration with Prof. C. Copéret (ETH Zürich)
25 MAS-DNP for studying protein-DNA interactions 10.0 kHz @ 14.1 T 395 GHz gyrotron Hp DnaB:ADP:ssDNA Measurement time 21 h. Lowest contour level: 2.1 times noise RMSD Morag, ..., Goldbourt, JACS , 2014 , 136 , 2292-2301; Wiegand, ..., Copéret, Böckmann, Meier, J. Biomol. NMR , submitted. Collaboration with Prof. C. Copéret (ETH Zürich)
26 Any chance for probing DNA interactions under conventional NMR conditions (in reasonable measurement time)? X CHHP & NHHP Morag, ..., Goldbourt, JACS , 2014 , 136 , 2292-2301.
27 CHHP 2D experiments for studying protein-DNA interactions Hp DnaB:ADP:AlF 4 - :ssDNA Measurement time 13 d. 200 μ s H-H spin diffusion time 11.74 T @ 17.0 kHz
28 Conclusions 31 P NMR allows to monitor nucleotide binding o 13 C/ 15 N CSPs highlight conformational o changes o Paramagnetic NMR allows to probe protein- ATP interactions o DNA binding can be detected in 31 P/ 13 C correlation experiments
Recommend
More recommend
Explore More Topics
Stay informed with curated content and fresh updates.