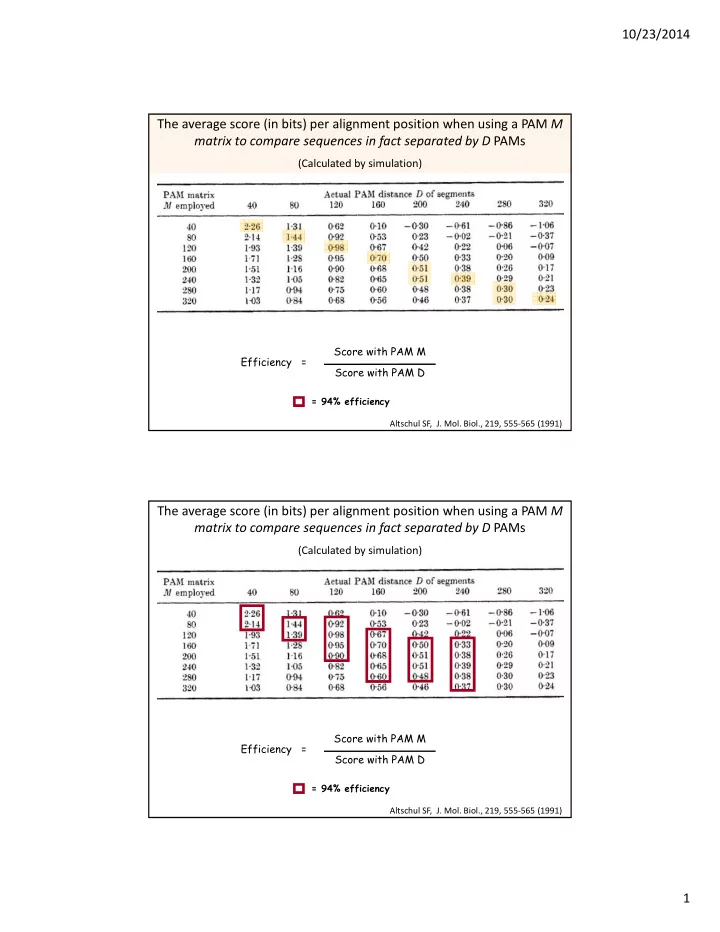
10/23/2014 1 The average score (in bits) per alignment position when using a PAM M matrix to compare sequences in fact separated by D PAMs
(Calculated by simulation)
Efficiency = Score with PAM M Score with PAM D
= 94% efficiency Altschul SF, J. Mol. Biol., 219, 555‐565 (1991)
The average score (in bits) per alignment position when using a PAM M matrix to compare sequences in fact separated by D PAMs
(Calculated by simulation)
Efficiency = Score with PAM M Score with PAM D
= 94% efficiency Altschul SF, J. Mol. Biol., 219, 555‐565 (1991)