1
Biochem ical Kinetics Made Easier
Petr Kuzmič, Ph.D.
BioKin, Ltd.1 . Theory: differential equations
- DYNAFI T software
2 . Exam ple I : Initial rate experiment
- p5 6 lck kinase / “ATP analog” inhibitor
3 . Exam ple I I : Time course experiment
- p3 8 α kinase / desatinib / competitive ligand displacement
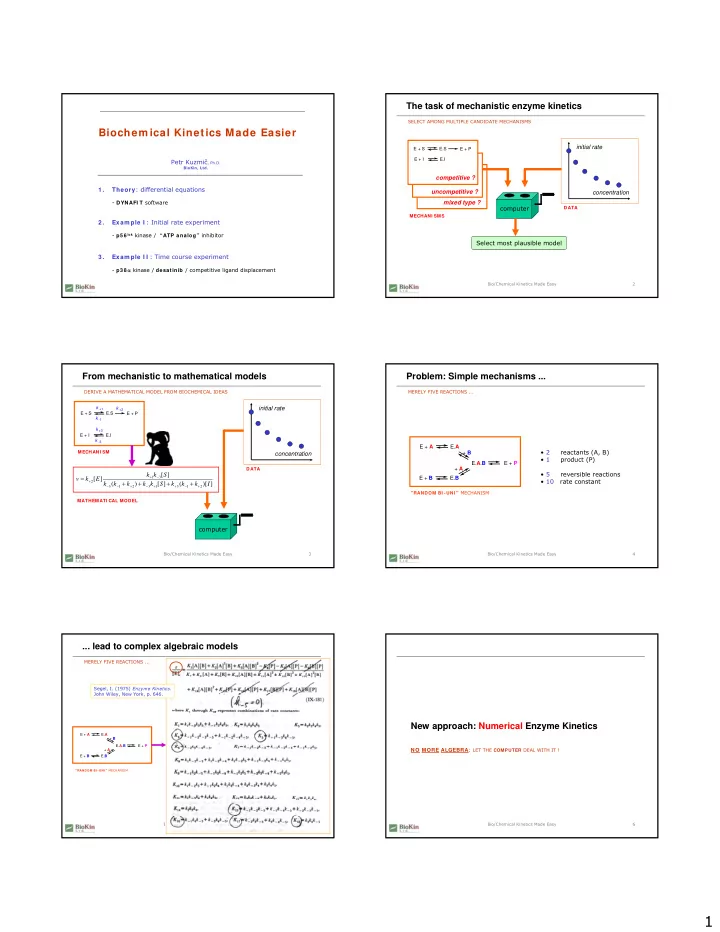
The task of mechanistic enzyme kinetics
SELECT AMONG MULTIPLE CANDIDATE MECHANISMSconcentration initial rate
DATAcomputer Select most plausible model
MECHANI SMScompetitive ?
E + S E.S E + P E + I E.Iuncompetitive ? mixed type ? competitive ?
Bio/Chemical Kinetics Made Easy 3From mechanistic to mathematical models
DERIVE A MATHEMATICAL MODEL FROM BIOCHEMICAL IDEASconcentration initial rate
DATAcomputer
MATHEMATI CAL MODEL E + S E.S E + P E + I E.I k +1 k -1 k +2 k +3 k -3] )[ ( ] [ ) ( ] [ ] [
2 1 3 1 3 2 1 3 3 1 2I k k k S k k k k k S k k E k v
+ − + + − + − − − + ++ + + + =
MECHANI SM Bio/Chemical Kinetics Made Easy 4Problem: Simple mechanisms ...
MERELY FIVE REACTIONS ...- 2
reactants (A, B)
- 1
product (P)
- 5
reversible reactions
- 10
rate constant
E + A E.A E + P E + B E.B E.A.B + B + A
"RANDOM BI -UNI " MECHANISM Bio/Chemical Kinetics Made Easy 5... lead to complex algebraic models
Segel, I. (1975) Enzyme Kinetics. John Wiley, New York, p. 646. E + A E.A E + P E + B E.B E.A.B + B + A "RANDOM BI - UNI " MECHANISM MERELY FIVE REACTIONS ... Bio/Chemical Kinetics Made Easy 6New approach: Numerical Enzyme Kinetics
NO MORE ALGEBRA: LET THE COMPUTER DEAL WITH IT !