SLIDE 3 23‐Mar‐15 3
Motifs in DNA sequences
- Inside gene coding regions: conserved genetic regions
- Outside coding regions: transcription factor binding sites
(TFBS)
– TATA box: found in promoters of Archaea and Eukaryotes, binds transcription factors or histones
Summarizing many aligned sequences
- Sometimes it is handy to summarize
hundreds of aligned sequences
- The consensus sequence is the sequence
containing the most frequent residues at each position
– The TATA box
- Unknown nucleotide: N
- Unknown nucleotide: N
- Unknown amino acid: X
Sequence profiles
- Consensus of a protein motif of 4 amino acids:
- A sequence profile shows much more detail:
Position 1 Position 2 Position 3 Position 4 A C 10 D 6 E F G H 7 I 7 K L M N P 2 Q R S 4 T V 1 W Y 3
Most conserved position
Sequence profiles “summarize” biochemistry
- A sequence profile represents all the possible sequences
and the sequence conservation at once
– Motifs – Protein families
- This is almost like describing the real
biochemical interactions!
...atG AA AT TT C ac... ...ccG AA GT TT C tg...
- We can predict that a conserved
position is important for the function
- f the protein because it is rarely
changed in evolution
– Which positions are more conserved – Which positions are less conserved
g ...agG AA AA TT C aa... ...gtG AA AT TT C cg... ...caG AA AT TT C tc... ...tgG AA AT TT C gt...
A 2 1 0 6 6 5 1 0 0 0 2 1 C 2 1 0 0 0 0 0 0 0 6 1 2 G 1 2 6 0 0 1 0 0 0 0 1 2 T 1 2 0 0 0 0 5 6 6 0 2 1
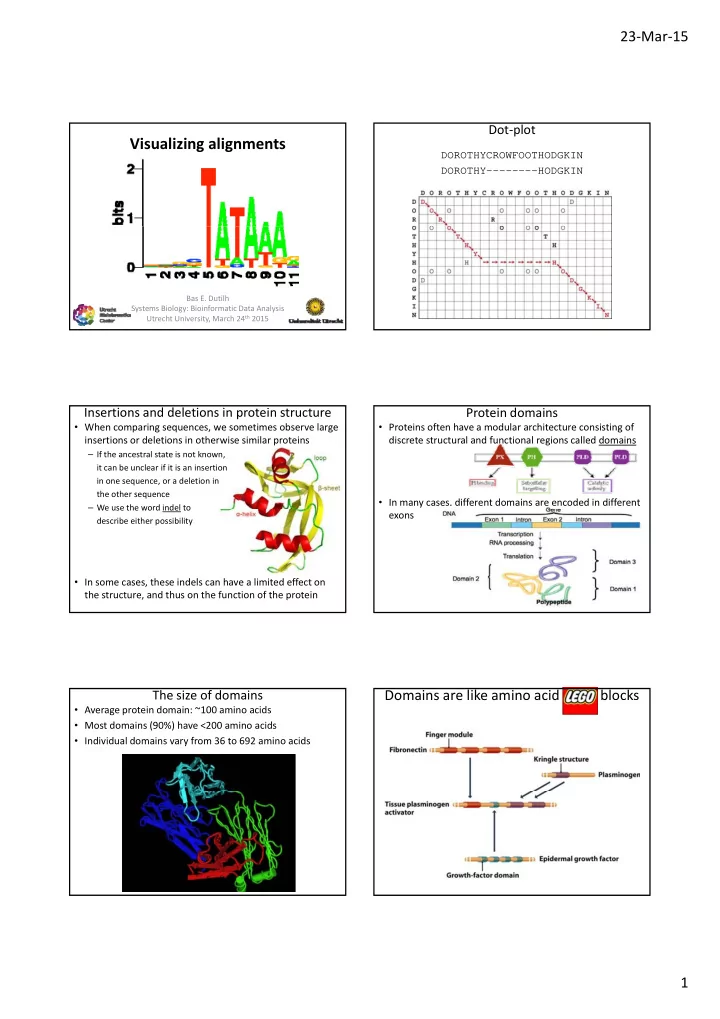
DNA sequence logos
- At each position, possible nucleotides are shown by
stacked letters
– Letter heights relative to frequencies pi (i = A, C, G, T)
- The total stack height shows the conservation
– Information content at position k (in bits):
) ( log ) 4 ( log ) (
2 , , , 2 i T G C A i i
p p k I
The TATA box
- Maximum information in a completely conserved position (e.g. always T)
– pA = 0; pC = 0; pG = 0; pT = 1 – Assume that 0 log2 (0) = 0 I = log2 (4) + (0 + 0 + 0 + 1 log2 (1)) = 2
- Minimum information in a completely unconserved position (random)
– pA = 0.25; pC = 0.25; pG = 0.25; pT = 0.25 I = 2 + (0.25 log2 (0.25) + 0.25 log2 (0.25) + 0.25 log2 (0.25) + 0.25 log2 (0.25)) I = 2 + (– 0.5 – 0.5 – 0.5 – 0.5) = 0
Transcription factor binding sites (TFBS)
- Transcription factors are proteins that bind a specific DNA
sequence