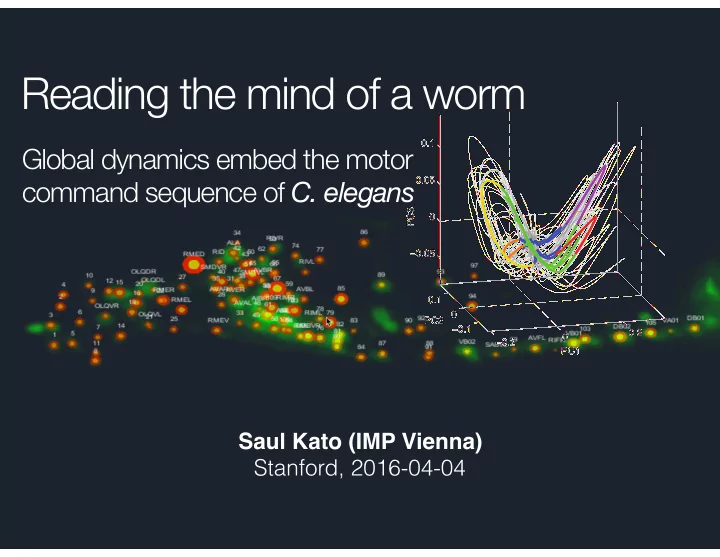
Reading the mind of a worm
Saul Kato (IMP Vienna) Stanford, 2016-04-04
−0.2 0.2 −0.1 0.1 −0.05 0.05 0.1 PC1 PC2 PC3
Reading the mind of a worm 0.1 Global dynamics embed the motor - - PowerPoint PPT Presentation
Reading the mind of a worm 0.1 Global dynamics embed the motor command sequence of C. elegans 0.05 PC3 0 0.05 0.1 0 PC2 0.1 0.2 0 0.2 PC1 Saul Kato (IMP Vienna) Stanford, 2016-04-04 collaborators Yifan Xu, Rockefeller Cori
−0.2 0.2 −0.1 0.1 −0.05 0.05 0.1 PC1 PC2 PC3
cannot rely on a homunculus or components
competent on a single trial basis behavior is an continuous,
system
Chemotaxis in E. coli Analysed by 3D Tracking Berg & Brown, Nature 1972
A0 = −kmA(1 − I) − kbA A + Kmm + kpGI
G0 = km(1 − I)A − kpIG + kr
Robustness and adaptation in simple biochemical networks Barkai & Leibler, Nature 1997
A0 = −kmA(1 − I) − kbA A + Kmm + kpGI
G0 = km(1 − I)A − kpIG + kr
Larry Abbott
302 neurons ~6000 synapses ~900 gap junctions
no classical action potentials. Na+ channels lost during nematode evolution
Virtual Worm Project
SLOWING DORSAL TURN FORWARD RUN VENTRAL TURN REVERSE RUN
interneurons
0.025 0.02 0.015 0.01 0.005 0.005 0.01 0.015 0.02 2 1.5 1 0.5 0.5 1 1.5 2 VB05 VB03 DD03 DD02 VB04 DD04 VD07 VB06 VD05 VD06 VD04 DB03 VC01 VC02 VA07 VD08 DB02 AS04 VA04 VD03 VA06 VC03 DB04 DD05 VA05 DA04 DA03 VD09 AS03 AS05 AS06 VB08 DD01 VD02 DA05 VA09 VA08 AS02 VA03 VB02 DB01 PDB VA02 VB09 VD10 PDA RID DA02 VB07 DA06 AS11 VD13 VD01 VA12 DD06 VC04 DA09 VD12 DVB PVDL AS09 VA11 DA08 DA07 VB11 AS08 PVDR VA10 PHCL PHBL VA01 AS01 VB10 AS07 DB07 PVCR PHBR AS10 PVCL AVAL DA01 LUAL DB05 DB06 AVAR PVWR PLML VD11 PHCR SABD LUAR PQR PVWL AVDL PVNR AVL AVDR AVBR PHAL AVBL PLMR PHAR FLPR AVG PVNL FLPL SABVR AVM SABVL AVJL PVPR DVC VC05 PVPL VB01 AVJR BDUR HSNR PDER PDEL PVM DVA AVFR RIFL AVFL RIFR AQR AVHR AVHL PVR SDQL ALMR BDUL AVKL ALA PVQR PVT ALML ADAL HSNL SDQR AVEL
normalized Laplacian eigenvector 2
ASJR ADER SAADL RMFR ADAR AVER ADLR SAAVR SAAVL ADLL AIML AVKR ASHR RIMR AIMR RIML ASHL PVQL SIBVL ASJL RMFL ASKR SAADR RIGL AIBR RICL RIS SMBVR AIBL PLNL ADEL ASKL PLNR RIGR SMBDL ALNL AIAR RICR SMBDR AIAL SMBVL RMGR ALNR AWBR RIR BAGR BAGL RMGL ASGL AUAL URXL ASGR AIZR AWBL SMDDR RIBL AIZL ASIR URYVL SIBVR URBL URYVR ADFR RMHL RIBR URYDR ASER AWAR ASIL AINR RIVL OLLL AUAR ADFL AWCR SMDDL RMHR RIVR SIBDL AWCL SIADL ASEL AINL AWAL URYDL CEPDL CEPVL OLLR RMDR URXR SIAVR CEPDR SIADR AIYL AIYR SIAVL SIBDR AFDR AFDL SMDVR SMDVL URBR RMDL CEPVR RIAL RIAR OLQDR OLQVL OLQDL RMDVR IL2L RMDDL RMED RMEV RIH RMDVL RMDDR IL1L IL2R IL1R URADL OLQVR IL1DL IL1VL RIPL IL1DR URADR RMEL IL1VR RIPR IL2DL URAVL IL2VL RMER IL2DR IL2VR URAVR
processing depth
interneurons sensory neurons motor neurons motor neurons sensory neurons Chen 2011 processing depth 2
CHEMOSENSORY NEURONS MECHANOSENSORY PHARYNX AMPHIDS (LR) INNER LABIA (6) OUTER LABIA (6) DEIRIDS HEAD (4) BODY TAIL PHASMIDS (LR) MOTONEURONS INTERNEURONS RING BODY
L RASE
L RASK
L RASJ
L RASG
L RAWC
L RAWB
L RURX
L R L RURY
L R L RURA
L R L RRME
L R L RSMD
L R L RSMB
L R L RSAA
L R L RSIB
L R L RSIA
L RFLP
L RBAG AQR
L RPHA
L RPHB
L RPHC
L RM2
L RM3
L RI2
L RI1
L RIL1
DL DR VL VRCEP
DL DR VL VROLQ
L ROLL
L RPDE
L RADE
L RPLM
L RPVD
L RALM
L RALN
L RAVM
L R L R L RIL2
L R L R L RAWA
L RLUA
L RBDU
L RPVQ
L RPVP
L RPVN
L RPVW
L RADE
L RASH
L RAVA
L RAVF
L RAVH
L RAVJ
L RAVK
L RRIA
L RRIF
L RRIG
L RRIC
L RRIB
L RURB
L RSDQ
L RAVB
L RPVC
L RAVD
L RAVE
L RADA
L RHSN AS 1 2 4 8 10 11 11
L RAIA
L RAIB
L RAIM
L RAIN
L RAIY
L RAUA
L RAIZ
L RRIM
L RRMF
L RRMH
L RRMG
L RRIP
L RRIV
L RAFD
L RASI
L RADF
L RADL PQR M1 M4 I3 I4 I5 I6 M5 MI MC NSM 3 5 6 9 7 VD 1 2 4 8 12 10 11 13 3 5 6 9 11 7 VB 1 2 4 8 10 11 11 3 5 6 9 7 VA 1 2 4 8 10 11 3 5 6 9 7 DA 1 2 4 8 9 3 5 6 7 DB 1 2 4 7 3 5 6 DD 1 2 4 3 5 6 VC 1 2 4 3 5 6 12
L RRMD
L R L R DSAB
D L D R awar awcr awcl avar aval avfr avfl avhr avhl avjr avjl avkr avkl riar riaL rifr rifl rigr rigl ricr ricL ribr ribL urbr urbl sdqr sdql avbr avbl pvcr pvcl avdr avdl aver avel adar adal hsnr hsnl as11 as01 vd13 vd01 vb11 vb01 va12 va01 da09 da01 db07 db01 dd06 dd01 vc06 vc01 aiar aial aibr aibl aimr aiml aimr aiml aiyr aiml auar aual aizr aizl rimr riml rmfr rmfl rmhr rmhl rmgr rmgl ripr ripl rivr rivl awal luar lual bdur bdul pvql pvqr pvpr pvpl pvnr pvnl pvwr pvwl ader adelAVG
avgALA
alaRID
ridRIR
rirRIS
risRIH
rihDVA
dvaDVC
dvcDVB
dvbPVM
pvmPDB
pdbPVR
pvrPVT
pvrPDA
pdaAVL
avlAVM
avm120 60 120 60 WT nlp-1 Odor Odor ΔF/F 200% time (s ) time (s)
GCaMP1.0
GCaMP3.0
trials
1 2
20 s
AWC
4 8 12 16 20 24
normalized magnitude lag (s)
y ( F/F) x
L N input
K(t) F(x) x
u(t) x(t) y(t)
input
kaf kas ka ks kf
dA dt = kaA+ input dS dt = ksS - k A =F S +
dF dt = kfF + kaf
as
A
10 20 30 40 Lag (s) 10 20 30 40 Lag (s)
F S
adapted from White, 1986
Retrovesicular ganglion Ventral ganglion Ventral ganglion Dorsal ganglion Anterior ganglion
10 µm
microfluidics spinning disk confocal 10 Z @ 2.9 volumes/s volumetric microscopy nuclear-localized GCaMP5K
M13$ GFP$ CaM$ NLS$ NLS$ N+$ +C$
AVAR AVAL RIMR RIML AVER VA01 SABVL
(OLQVR/URYVR) (OLQDL/URYDL) AIBL AIBR 42 108 38 (OLQDR/URYDR) 9 20 93 40 ALA 31 47 22 50 AVFL 48 76 18 55 52 77 94 62 90 5 85 49 10 14 1 11 (RIFR/AVG/DD01) 51 88 83 35 105 (SMBDR/SMBVR/SMDDR/RMFR) 64 74 7 25 34 86 3 6
60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020 1080
Time (s)
Neuron Time (s) 60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020 ΔF/F0 −0.2 0 1 1.5 AVAR AVAL RIMR RIML AVER VA01 SABVL OLQVL DB01 VB01 DB02 RMER RMEL RID AVBR RIBL VB02 RMED RMEV AVBL SMDVL SMDVR RIVL RIVR AIBL OLQVR AIBR OLQDL OLQDR RIFR SMBVR PC1 PC2 PC3 60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020 3 2 1 Time (s) TPC#
60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020 3 2 1 Time (s) TPC#
12345678910 20 40 60 80 PC Variance explained (%)
60 120 180 240 300 360 420 480 540 600 660 3 2 1 Time (s) TPC#
60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020 3 2 1 Time (s) TPC#
−0.2 0.2 −0.1 0.1 −0.05 0.05 0.1 PC1 PC2 PC3
AVAL ↑ SMDV ↑ RMED ↑
0.2 −0.1 0.1 PC1 PC2 −0.2
RIVL RIVR RMED RMEL RMER RMEV SABVL (SMBDR/SMBVR/ SMDDR/RMFR) SMDVL SMDVR VA01 VB01 VB02 AIBL AIBR ALA AVAL AVAR AVBL AVBR AVER AVFL DB01 DB02 (OLQDL/ URYDL) (OLQDR/ URYDR) (OLQVL/ URYVL) (OLQVR/ URYVR) RIBL RID (RIFR/AVG/ DD01) RIML RIMR
30 Hz epifluorescence 10 Hz infrared
Faumont et al, 2011
∆ R / R0 1 1.5 2 2.5 1 2 3 4 1 2 3 4 Time from reversal start (s) 1 2 ∆ R / R0 ∆ R / R0 ∆ R / R0
1 2
AVA RIM AVE AIB
Time from reversal end (s)
1 2 .5 1 −1 1 −1 1 1
REVERSAL
n>100
FALL HIGH RISE LOW
π 3π π 2 2 RISE HIGH FALL LOW
60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020 3 2 1 Time (s) TPC#
FALL HIGH RISE LOW
π 3π π 2 2 RISE HIGH FALL LOW
60 s
π 3π π 2 2 RISE HIGH FALL LOW
AIBL SMDVL SABVL
AVAR AVAL RIMR RIML AVER VA01 SABVL AIBL AIBR
60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020
OLQVL
high-to-fall transition timing
1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 AIBL AIBR AVAL AVAR AVER OLQDL OLQDR OLQVL OLQVR RIML RIMR SABVL SMDVL VA01 avbl avbr avfl db01 db02 ribl rid rifr rivl rivr rmed rmel rmer rmev smbvr smdvr vb01 vb02
t=0 none
+6
seconds
π 3π π 2 2 RISE HIGH FALL LOW
1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 AIBL AIBR AVAL AVAR AVER OLQDL OLQDR OLQVL OLQVR RIML RIMR SABVL SMDVL VA01 avbl avbr avfl db01 db02 ribl rid rifr rivl rivr rmed rmel rmer rmev smbvr smdvr vb01 vb02
FALL2 FALL1
−0.2 −0.15 −0.1 −0.05 0.05 0.1 0.15 0.2 0.25 0.3 −0.15 −0.1 −0.05 0.05 0.1 0.15 −0.05 0.05 0.1 PC1 PC2 PC3FALL1 FALL2
1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 AIBL AIBR AVAL AVAR AVER OLQDL OLQDR OLQVL OLQVR RIML RIMR SABVL SMDVL VA01 avbl avbr avfl db01 db02 ribl rid rifr rivl rivr rmed rmel rmer rmev smbvr smdvr vb01 vb02
FALL2 FALL1
1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 2 AIBL AIBR AVAL AVAR AVER OLQDL OLQDR OLQVL OLQVR RIML RIMR SABVL SMDVL VA01 avbl avbr avfl db01 db02 ribl rid rifr rivl rivr rmed rmel rmer rmev smbvr smdvr vb01 vb02
FALL2 FALL1
GCaMP/mCherry 0.5 ∆ R/R0 head-bend angle 30°
SMDV
V D Time (seconds) 30 60 90
D V REV
Free moving imaging
∆ R / R0 1 1.5 2 2.5 1 2 3 4 1 2 3 4 Time from reversal start (s) 1 2 ∆ R / R0 ∆ R / R0 ∆ R / R0
1 2
AVA RIM AVE AIB
Time from reversal end (s)
1 2 .5 1 −1 1 −1 1 1
REVERSAL
n>100
Time from end (s)
1 2 Time from end (s)
1 2 Ventral Dorsal Reverse Reverse
SMDV
TURNS
PC1
0.2
PC2
0.1
SUSTAINED REVERSAL REVERSAL II REVERSAL I VENTRAL TURN DORSAL TURN FORWARD SLOWING
PC1
0.2
PC2
0.1
REVS REV1 REV
2
DORSAL TURN VENTRAL TURN SLOWING FWD
BRAIN STATE GRAPH
0.2
0.1
PC1 PC2
BRAIN STATE GRAPH
REVS REV1 REV
2
DORSAL TURN VENTRAL TURN SLOWING FWD
BEHAVIORAL STATE
SLOWING DORSAL TURN FORWARD RUN VENTRAL TURN REVERSE RUN
REVERSAL DORSAL TURN VENTRAL TURN SLOWING FWD
60 120 180 240 300 360 420 480 540 600 660 720 780 840 900 960 1020 3 2 1 Time (s) TPC#
−0.2 −0.1 0.1 −0.15 −0.1 −0.05 0.05 0.1 0.15 PC1 PC2
Pokala, PNAS 2014
−0.2 0.2 0.4 −0.2 −0.1 0.1 −0.1 0.1 PC1 PC2 PC3
Fall Rise
4% O 21% O BAG
2 2
Pre
Reversal comand state probability Time (s) O2 4% 4% 4% 4% 4% 4% 21% 60 120 180 240 300 360 420 480 540 600 660 720 0.25 0.5 0.75 1
n=13
scaffold for detailed studies
input
−5
−4
−3
−2
−1
Clustering p-value
π 3π π 2 2 RISE HIGH FALL LOW
−3π −2π −π π 10
−5
10
−4
10
−3
10
−2
10
−1
10 Global phase Fall sub-trajectories −2π −π π 2π 10
−5
10
−4
10
−3
10
−2
10
−1
10 p−value Clustering Global phase Rise sub-trajectories
π 3π π 2 2 RISE HIGH FALL LOW
REVS REV1 REV
2DORSAL TURN VENTRAL TURN SLOWING FWD
0.2
PC1
0.1 0.1
PC2 PC3 PC1
0.2
PC2
0.1 drive (a.u.) no value
0.5 1
2 4 6 8 10 12 14 16 18 20 >20 0.5 LOW phase probability 2 4 6 8 10 12 14 16 18 20 >20 0.2 0.4 RISE phase probability 2 4 6 8 10 12 14 16 18 20 >20 0.05 0.1 HIGH phase probability 2 4 6 8 10 12 14 16 18 20 >20 0.1 0.2 FALL phase probability state duration (s)
time (s)
10 20 30 40 50 60 70 80 90 100 110 120 130 140 150 160 170
Vm
Mag (a.u.)
0.5 1
time (s)
30 60 90 120 150 180 210 240 270 300
Vm
Mag (a.u.)
0.5 1
time (s)
30 60 90 120 150 180 210 240 270 300 330 360 390
Vm
Mag (a.u.)
0.5 1
time (s)
30 60 90 120 150 180 210 240 270
Vm
Mag (a.u.)
0.5 1
c
Mark Alkema