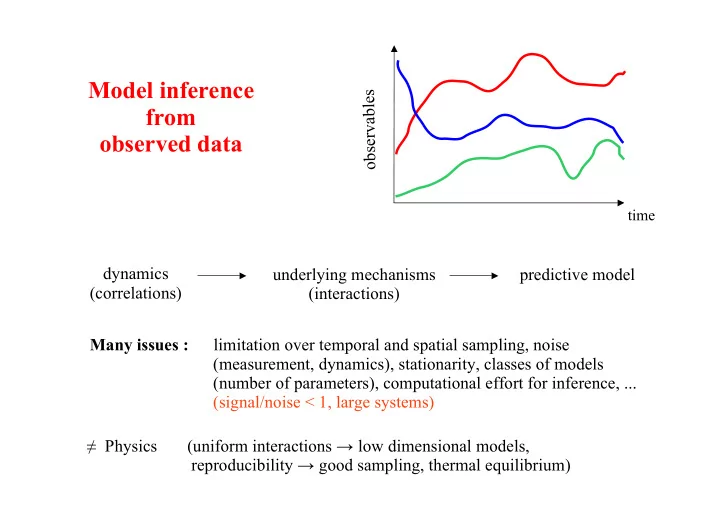
Model inference from
- bserved data
time
dynamics (correlations) underlying mechanisms (interactions)
- b
s e r v a b l e s predictive model Many issues : limitation over temporal and spatial sampling, noise (measurement, dynamics), stationarity, classes of models (number of parameters), computational effort for inference, ... (signal/noise < 1, large systems) ! Physics (uniform interactions " low dimensional models, reproducibility " good sampling, thermal equilibrium)