Gradient Analysis
Multivariate Fundamentals: Rotation/Distance
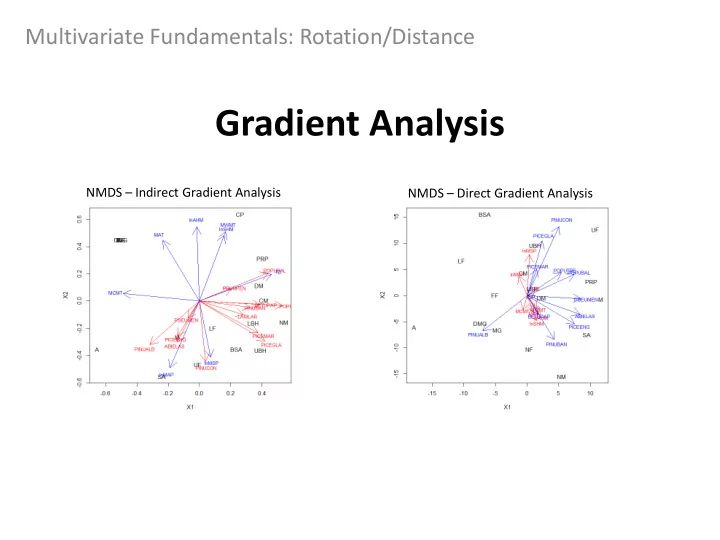
NMDS – Indirect Gradient Analysis NMDS – Direct Gradient Analysis
Gradient Analysis NMDS Indirect Gradient Analysis NMDS Direct - - PowerPoint PPT Presentation
Multivariate Fundamentals: Rotation/Distance Gradient Analysis NMDS Indirect Gradient Analysis NMDS Direct Gradient Analysis Objective: Use one dataset to explain another Use the spatial patterns of each dataset to try and understand the
NMDS – Indirect Gradient Analysis NMDS – Direct Gradient Analysis
E.g. environmental factors, species populations and characteristics of communities
MAT MAP AHI … Sp1 Sp2 Sp3 …
Predictor Variables Response Variables
AHI MAP MAT Sp3 Sp1 Sp2 Example: Species 1 and 2 are associated with greater MAT and less MAP (warm & dry)
AHI MAP MAT Sp3 Sp1 Sp2
Example: Species 1 and 2 are associated with moderate MAT and moderate MAP (mild & moist)
Can extend any ordination technique to a gradient analysis But the easiest way to look at both a direct and indirect gradient analysis is to use a PCA (rotation) or NMDS (distance) plot and simply add a second set of vectors to infer relationships between datasets (we do this in Lab 7)
NMDS – Indirect Gradient Analysis NMDS – Direct Gradient Analysis
Indirect: Lodgepole pine (PINUCON) is associated with environments with greater growing season precipitation (lnMSP) Direct: Lodgepole pine is still associated with wetter summer environments, but so is white spruce (PICEGLA)
Indirect gradient analysis Direct gradient analysis
E.g. brings out pairs of variables between datasets that are highly associated with each other
Rotation based technique for constrained gradient analysis Rotate both the predictor and response datasets independently to maximize correlation between corresponding variables among datasets Once correlation is maximized between datasets on the first axes – the axis for each dataset is fixed and rotation for the second predictor/response variable is carried out Repeated for all variables You do not need to have the same number of response and predictor variables Conical functions (rotations) will be built for the smaller number of variables Herold Hotelling (1895-1974)
Predictor Predictor Predictor Response Response Response
Golden Can fit linear curve BUT hard to fit – should transform response or predictor variables CANCOR (and others) will fail to detect a relationship Use MRT (Lab 8)
CANCOR in R:
library(CCA) cancor(predictorData,responseData) (CCA package) cc(predictorData,responseData)
E.g. Environmental Variables
E.g. Species Community Variables
The cancor() function will display the correlation values The cc () function outputs a number of statistics that we can query to provide some more information from the analysis AND use to test significance Predictor data table and Response data table need to have the same number of rows BUT they do not need to have the same number of columns
Correlations between the rotated axes The first correlation will be the maximum values and all successional correlations will be smaller Estimates of the predictor and response coefficients from the rotation model (matrix algebra) The value used to adjust each predictor and response variable under rotation
We can use the output from the cc
function to individually test if the
correlation values between our axes are significant P-values test the hypothesis that the true correlation is equal to 0 i.e. There is no correlation Therefore small p-values reject this hypothesis and there is a true significant correlation between the axes Based on our CANCOR analysis of 3 predictor and 3 response variables, the correlations found between rotation 1 (0.93) and rotation 2 (0.7) were found to be significant BUT the correlation between rotations 3 (0.12) was found to NOT be significant
If correlation for the canonical functions (rotated axes) is significant we can look at the loadings to see what each new rotated axes is related to in our original predictor and response variables In our case we only have to look at Can1 and Can2 We now have to interpret the loading values for the predictor and response variables together (e.g. associate high loadings together) Can1: When Env3 is rare, Spec 2 and Spec 3 have lower frequency (both negative) Spec2 and Spec3 prefer Env3 (reverse) Can2: When Env1 is abundant, Spec1 has a higher frequency (both positive) Spec 1 prefers Env1 Nothing really likes Env2
Rotation-based technique for constrained gradient analysis Like CANCOR, CCA aims to maximize the correlations between response and predictor variables, BUT response scores are constrained to be linear combinations of predictor variables in a effort to maximize the variance explained by the predictor data in all
Multiple linear regression is used to solve the linear combinations of predictor variables Categorical variables can be used in CCA – converted to “dummy” variables where each class is assigned a numeric value (should be addressed with caution) CCA is considered an improvement over CANCOR in some fields CCA was developed for Ecology Like CANCOR – linearity of the relationship between response and predictor variables is assumed CCA may be able to detect some non-linear responses, BUT there are better techniques for that (MRT, RandomForest – Lab 8)
CCA in R:
library(vegan) cca(responseData,Predictor1+Predictor2+…,data=predictorData) (vegan package)
E.g. Species Community Variables
A linear equation including the predictor variables (e.g. Environmental Variables) that you feel are related to the response variable outputs (e.g. Species Occurrence) You can include as many predictors as you wish HOWEVER, the more predictors you include the more complex the analysis and the capacity to detect strong relationships is reduced (so pick your predictor variables mindfully)
E.g. Environmental Variables
Predictor data table and Response data table need to have the same number of rows BUT they do not need to have the same number of columns
We analyzed a model where we included 3 environmental variables to explain species frequency Variance Explained: Total variance – total amount of variance in the response variables (e.g. species data) Constrained variance – how much is explained by the predictor variables (e.g. environmental data) Unconstrained variance – how much variance is left in the response variables (unexplained) Eigenvalues – how much of the variance is explained by the individual axes of the
You will have to figure out the % of variance explained by yourself Simply divide value/ total variance Constrained CCA1 = 0.005395/0.007953 = ~68% Constrained CCA2 = 0.000214/0.007953 = ~3%
Unconstrained defaults to a correspondence analysis (unconstrained)
To determine if a significant relationship between our response and predictor variables exists we can run our CCA output through an ANVOA Generic ANOVA tells us if a significant relationship between the response and predictor variables exists ANOVA (overwrites as anova.cca in vegan package) For by option – selecting "term" p-values will be produces for each predictor term For permu option – the number of permutations to use to generate the p-values P-values test the hypothesis that the correlation between species variables and each environmental variable is 0 From p-values Env2 and Env3 are significantly associated with species occurrences
From the image we can see:
Env3 appears positively associated with Spec3 and negatively associated with Spec2 and Spec1 Env2 appears positively associated with Spec3 and Spec1 and negatively associated with Spec2 Env1 appears negatively associated with Spec2 – but from ANOVA this is not a significant relationship
Rotation-based technique for constrained gradient analysis The goal of RDA is to apply linear regression in order to find linear combinations of predictor variables to represent as much variance in the response variables as possible CCA focuses more on species composition, i.e. relative abundance, if you have a gradient along which all species are positively correlated, RDA will detect such a gradient while CCA will not With RDA, it is possible to use 'species' that are measured in different units, BUT in this case, the data must be centered and standardized RDA can useful when gradients are short or you are conducting a short-term experimental study Like CCA categorical variables can be used in RDA – converted to “dummy” variables where each class is assigned a numeric value (should be addressed with caution) Like CANCOR and CCA – linearity of the relationship between response and predictor variables is assumed
RDA in R:
library(vegan) rda(responseData,Predictor1+Predictor2+…,data=predictorData) (vegan package)
E.g. Species Community Variables
A linear equation including the predictor variables (e.g. Environmental Variables) that you feel are related to the response variable outputs (e.g. Species Occurrence)
E.g. Environmental Variables
Predictor data table and Response data table need to have the same number of rows BUT they do not need to have the same number of columns With RDA it is possible to use response variables that are measured in different units BUT in this case the dependent data must be centered and standardized before executing the analysis to do this you can specify the option scale=TRUE (default is FALSE)
We analyzed a model where we included 4 environmental variables to explain species frequency for 6 species Eigenvalues – how much of the variance is explained by the individual axes of the
You will have to figure out the % of variance explained by yourself Simply divide value/ total variance Constrained RDA1 = 74.52/112.88889 = ~66% Constrained RDA2 = 24.94/112.88889 = ~22% Constrained RDA3 = 8.88/112.88889 = ~8% Variance Explained: Total variance – total amount of variance in the response variables (e.g. species data) Constrained variance – how much is explained by the predictor variables (e.g. environmental data) Unconstrained variance – how much variance is left in the response variables (unexplained)
Unconstrained defaults to a PCA analysis (unconstrained)
To determine if a significant relationship between our response and predictor variables exists we can run our RDA output through an ANVOA (like CCA) Generic ANOVA tells us if a significant relationship between the response and predictor variables exists ANOVA (overwrites as anova.cca in vegan package) For by option – selecting "term" p- values will be produces for each predictor term For permu option – the number of permutations to use to generate the p- values P-values test the hypothesis that the correlation between species variables and each environmental variable is 0 From p-values Depth, Sand, and Coral are all significantly associated with fish occurrences “Other substrate” was removed from the analysis due to collinearity (last column entered)
From the image we can see:
All species were found to dislike environments characterized by Sand (i.e all species respond the same across an environmental gradient) Sp3 and Sp4 associated with Coral environments Sp1 and Sp2 associated with environments with greater water Depth