SLIDE 1
CSE 421 Midterm Scores
1
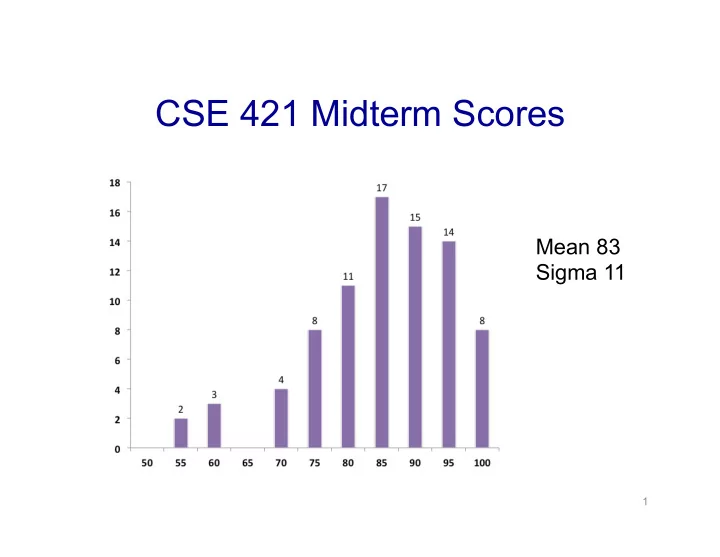
CSE 421 Midterm Scores Mean 83 Sigma 11 1 CSE 421 Algorithms - - PowerPoint PPT Presentation
CSE 421 Midterm Scores Mean 83 Sigma 11 1 CSE 421 Algorithms Sequence Alignment 1 Sequence Alignment Goal: position characters in strings so they best line up with one another We can do this via Dynamic Programming 2 What is an
1
1
2
3
4
5
6
1.
2.
7
8
9
10
11
12
13
14
T
A
2 1
C
1 4 3
G
3
★ T
…
…
T
T à S v
15
T
A
2 1
C
1 4 3
G
3
★ T
…
…
T
T à S v
16
T
A
2 1
C
1 4 3
G
3
★ T
…
…
T
T à S v
17
C G T … T
A
2 1
C
1 4 3
G
3
★ T
…
…
T
`8
T à S v
19
T à S v
20
T à S v
21
T à S v
22
T à S v
23
T à S v
24
Corresponding Alignment: CATGT
21
25
26
27
28
29