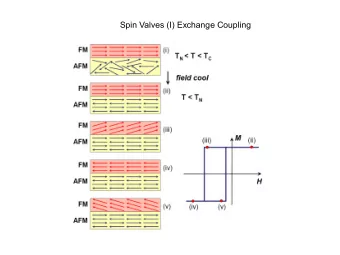
Coupling ReaxFF and DREIDING to Coupling ReaxFF and DREIDING to - PowerPoint PPT Presentation
Coupling ReaxFF and DREIDING to Coupling ReaxFF and DREIDING to Model Enzymatic Reactions Model Enzymatic Reactions Li Tao, Markus J. Buehler and Li Tao, Markus J. Buehler and William A. Goddard William A. Goddard Motivation Motivation
Coupling ReaxFF and DREIDING to Coupling ReaxFF and DREIDING to Model Enzymatic Reactions Model Enzymatic Reactions Li Tao, Markus J. Buehler and Li Tao, Markus J. Buehler and William A. Goddard William A. Goddard
Motivation Motivation � Find efficient computational method to model Find efficient computational method to model � reactivity in large biological systems reactivity in large biological systems � Existing QM/MM methods can model only a few Existing QM/MM methods can model only a few � pre- -selected atoms selected atoms pre • Enzymatic reactions may involve hundreds or Enzymatic reactions may involve hundreds or • thousands of reactive atoms thousands of reactive atoms • Not feasible with QM/MM schemes Not feasible with QM/MM schemes • � ReaxFF can model much larger regions ReaxFF can model much larger regions � involving several thousands of atoms. involving several thousands of atoms. • not practical for entire biological system not practical for entire biological system • � 80,000 iterations on 280 80,000 iterations on 280- -atom system atom system � • • DREIDING DREIDING – – dynamics took 1h 41m dynamics took 1h 41m • ReaxFF ReaxFF – – dynamics took 10h 26m dynamics took 10h 26m •
Comparison MM, ReaxFF, QM Comparison MM, ReaxFF, QM Estimated Clocktime Maximum Able to for 1 ns number of Model (max. # atoms Reactivity atoms) MM (DREIDING) 100,000 2 days no ReaxFF 3,000 5 days yes months, QM (DFT) 500 years yes Compromise: Hybrid ReaxFF/ MM scheme Compromise: Hybrid ReaxFF/ MM scheme Allows: Large systems (> 100,000 atoms Allows: Large systems (> 100,000 atoms with ~ 3,000 reactive atoms) with ~ 3,000 reactive atoms)
Coupled ReaxFF and DREIDING Coupled ReaxFF and DREIDING � Previous ReaxFF studies Previous ReaxFF studies Other regions � important too? on enzymes (subtilisin subtilisin, , on enzymes ( lysozyme) fixed non- - lysozyme) fixed non Localized reaction zone participating atoms participating atoms Enzyme � This region is important This region is important � • Elasticity, conformational Elasticity, conformational • changes, inhibitors changes, inhibitors � Treating non Treating non- -active active � region with DREIDING region with DREIDING allows physical forces to allows physical forces to be modeled be modeled Active Substrate Site
Implementation – – Coupling of force Coupling of force Implementation fields using mixed Hamiltonians fields using mixed Hamiltonians Energy Calculation: 100% 50% ReaxFF, 100% DREIDING 50% DREIDING ReaxFF Calculated with ReaxFF Ghost atoms (0% ReaxFF) Interpolation linearly or smoothly using sine function � CMDF framework allows to assign weights CMDF framework allows to assign weights � � Use transition zone of radius Use transition zone of radius r r t � t � “Ghost atoms” “Ghost atoms” �
The ModMulti ModMulti Modules Modules The New CMDF Modules for code coupling New CMDF Modules for code coupling � ModMulti ModMulti � • Functions for selecting atoms, assigning Functions for selecting atoms, assigning • weights (linearly and non- -linearly) linearly) weights (linearly and non � ModRestraints ModRestraints � • Functions for driving reactions using restraints Functions for driving reactions using restraints • (see next slide) (see next slide) � OBtools OBtools � • Utility functions • Utility functions • File output in BGF format File output in BGF format • • Manipulating X • Manipulating X OpenBabel OpenBabel objects objects
Implementation - - Coupling Coupling Implementation � Regions selected Regions selected � using python using python assignsphere_ w eights( ) function functions functions Ghost atoms � Overlapping Overlapping � weights for bigger Transition weights for bigger Region regions regions Pure ReaxFF � Regions can be Regions can be � reassigned as reassigned as reaction progresses reaction progresses Pure DREIDING
Implementation - - Restraints Restraints Implementation Why - Chemical reactions occur slowly at room temperature (biology) MD currently limited to nanoseconds – Chemical reactions need to be assisted to overcome barrier r A Bond Restraint Energy Barrier ) ( B ( ) v Bond restraints – – keep distances keep distances Bond restraints − 2 = − k r r � � f k 1 e 2 eq u between two atoms at specified between two atoms at specified 1 equilibrium distance equilibrium distance • • Equilibrium distance can change Equilibrium distance can change linearly over time to drive reactions linearly over time to drive reactions Angle restraints – Angle restraints – control angle control angle � � θ between three atoms between three atoms Angular Restraint
Wave propagation Wave propagation � Apply sudden jolt to Apply sudden jolt to Strain on each carbon � 2.0 ps end of C 80 H 162 chain end of C 80 H 162 chain � Wave propagated Wave propagated � through ReaxFF region through ReaxFF region � Demonstrates Demonstrates Pure Reax � seamless coupling 4.0 ps seamless coupling Transition Ghost 6.0 ps
Single molecule tensile test: Single molecule tensile test: Stretching a C 80 H 162 chain Stretching a C 80 H 162 chain Atomistic model: F F ReaxFF � Apply forces to a hydrocarbon chain to investigate Apply forces to a hydrocarbon chain to investigate � how the chain breaks how the chain breaks • Relationship between temperature and breaking strain Relationship between temperature and breaking strain • � Ensure coupling is done correctly Ensure coupling is done correctly � � Same weights as before: 15 carbon atoms in Same weights as before: 15 carbon atoms in Reax Reax, , � 10 A transition zone 10 A transition zone Transition and Ghost Pure ReaxFF Transition Pure DREIDING and Ghost
Results Results Temperature vs. Breaking Strain 1.28 1.26 Breaking Strain 1.24 1.22 1.2 1.18 1.16 1.14 1.12 1.1 1.08 0 100 200 300 400 500 600 700 Temperature (K) � Strain for breakage decreases with Strain for breakage decreases with � temperature temperature
Modeling a Simple Reaction Modeling a Simple Reaction System Energy (kcal/mol) O O H H C 80 H 161 O Transition State O O H H O - C 80 H 161 ~10kcal End State O O H H C 80 H 161 O Initial State Number of steps • System energy • Moving Average
Summary Summary � Have achieved coupling of ReaxFF and Have achieved coupling of ReaxFF and � DREIDING DREIDING � Demonstrated coupling by propagating Demonstrated coupling by propagating � waves through the molecule waves through the molecule � Applied this method to modeling breaking Applied this method to modeling breaking � strain of single C 80 H 162 molecule as a strain of single C 80 H 162 molecule as a function of temperature. function of temperature. � New method allows coupling ~ 3,000 New method allows coupling ~ 3,000 � reactive atoms with 100,000 nonreactive reactive atoms with 100,000 nonreactive atoms atoms
Modeling Enzymatic Modeling Enzymatic Activity of Subtilisin Subtilisin Activity of � Test coupling of force fields Test coupling of force fields � on biological system on biological system � Subtilisin Subtilisin is a serine protease is a serine protease � from bacteria from bacteria � Active site consists of Active site consists of � catalytic triad (Ser, His, Asp) catalytic triad (Ser, His, Asp) � Entire system of 4,000 atoms Entire system of 4,000 atoms � • Too large for pure Too large for pure ReaxFF ReaxFF • Number of atoms treated by Number of atoms treated by ReaxFF: 1210 (ca. 30%) ReaxFF: 1210 (ca. 30%) Entire: 3933 Entire: 3933
Procedure for Modeling Enzymatic Procedure for Modeling Enzymatic Activity of Subtilisin Subtilisin Activity of Minimize energy, then equilibriate equilibriate system at 300 K system at 300 K Minimize energy, then • • Model each reaction step using restraints to drive. Model each reaction step using restraints to drive. • • • Our case: First step • Our case: First step – – Transfer proton from serine to Transfer proton from serine to histidine histidine Compare energy barriers with pure ReaxFF and QM results Compare energy barriers with pure ReaxFF and QM results • • Asp Energy Moving average (150 points) O -77125 O - -77150 H Energy (kcal/mol) -77175 His N -77200 H O NH -77225 N H H H H H -77250 N O -77275 O H 2 C -77300 O 2 N Step1 Step2 Step3 Step4 Step5 Step6 Ser Step 1 – Proton Transfer Reaction coordinate (Pure ReaxFF)
Conclusion and Outlook Conclusion and Outlook � Possible alternative to QM/MM methods, Possible alternative to QM/MM methods, � but simpler to use and much faster but simpler to use and much faster � Coupled calculations are more efficient Coupled calculations are more efficient � than pure ReaxFF (tradeoff) than pure ReaxFF (tradeoff) � Possibly useful for quick scanning of Possibly useful for quick scanning of � reaction pathways reaction pathways • Designing enzyme with improved enzymatic Designing enzyme with improved enzymatic • activity activity • Understanding biological mechanisms Understanding biological mechanisms •
Recommend
More recommend
Explore More Topics
Stay informed with curated content and fresh updates.