INSTITUTE OF THEORETICAL INFORMATICS – ALGORITHMICS
COBS: A Compact Bit-Sliced Signature Index
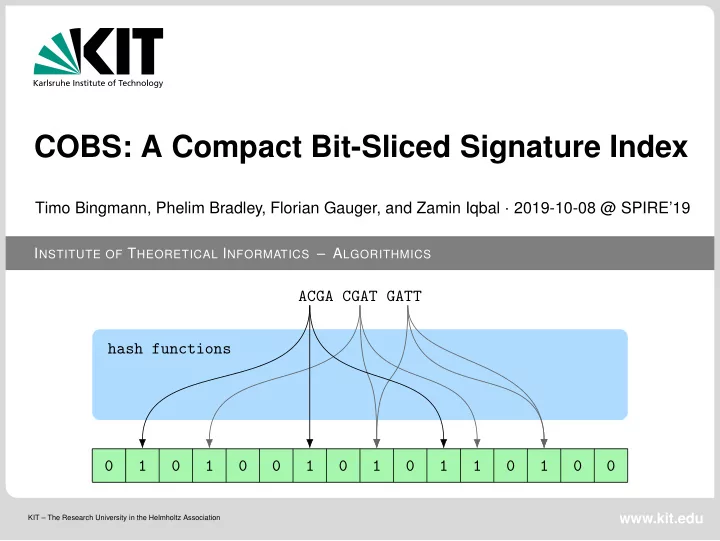
Timo Bingmann, Phelim Bradley, Florian Gauger, and Zamin Iqbal · 2019-10-08 @ SPIRE’19 ACGA CGAT GATT hash functions 1 1 1 1 1 1 1
KIT – The Research University in the Helmholtz Association
www.kit.edu