SLIDE 1 Analysis of variance (ANOVA)
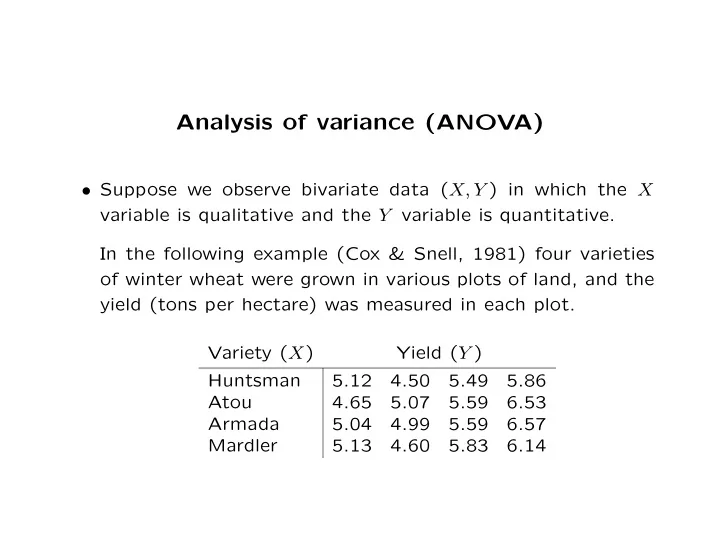
- Suppose we observe bivariate data (X, Y ) in which the X
variable is qualitative and the Y variable is quantitative. In the following example (Cox & Snell, 1981) four varieties
- f winter wheat were grown in various plots of land, and the
yield (tons per hectare) was measured in each plot. Variety (X) Yield (Y ) Huntsman 5.12 4.50 5.49 5.86 Atou 4.65 5.07 5.59 6.53 Armada 5.04 4.99 5.59 6.57 Mardler 5.13 4.60 5.83 6.14

SLIDE 2
The X variable is the type of wheat, which is qualitative, and in this context is called a factor. Specifically, it is a four level factor, since there are four types of wheat. In general, we will use m to denote the number of levels of the factor. All 16 data values are assumed to be independent. The four values in a given row are independent and identically distributed (iid), and are referred to as replicates. Note that this implies a key assumption – it is assumed that the mean and variance within each row are fixed. Our primary interest will be whether the means for different rows (different varieties of wheat) differ. This would imply that some varieties of wheat are better than others. The analysis is easiest when we assume that the variances for all rows are the same.

SLIDE 3 This type of data is called a balanced one-way layout. The term “balance” refers to the fact that there are the same number of observations in every row. The term “one-way” refers to the fact that there is only one X variable.
- Our notation for this type of data will be Yij, where i =
1, 2, 3, 4 indicates the type of wheat (i.e. 1 = Huntsman, 2 = Atou, 3 = Armada, 4 = Mardler), and j indexes the
- replicates. Thus, Y11 = 5.12, Y12 = 4.50, Y21 = 4.65, Y44 =
6.14, etc. Additional notation: Yi· =
j Yij is the sum of all values
in the ith row, n is the number of values in each row (4 in the example above), ¯ Yi· = Yi·/n is the average value in the ith row, Y·· =
ij Yij is the sum of all observations, and
¯ Y·· = Y··/mn is the overall (“grand”) mean.

SLIDE 4 All of these values can be displayed in an ANOVA table: Variety (X) Yield (Y ) Yi· ¯ Yi· Huntsman 5.12 4.50 5.49 5.86 20.97 5.24 Atou 4.65 5.07 5.59 6.53 21.84 5.46 Armada 5.04 4.99 5.59 6.57 22.19 5.55 Mardler 5.13 4.60 5.83 6.14 21.70 5.43 86.7 5.42 where the values in the lower right are Y·· = 86.7 and ¯ Y·· = 5.42.
- Analysis of variance (ANOVA) specifies the following simple
model for these data:

SLIDE 5
Yij = µ + αi + ǫij. The constant values µ, α1, α2, α3, and α4 are unknown pa- rameters of the population, and the random variables ǫij are iid errors with mean 0 and a common unknown variance σ2. For example, the model gives the following for certain specific data points: Y11 = µ + α1 + ǫ11 Y14 = µ + α1 + ǫ14 Y23 = µ + α2 + ǫ23 Y42 = µ + α4 + ǫ42.

SLIDE 6
- One difficulty with the above model is that different values
- f µ and the αi will give the same mean values for every data
point. Specifically, if we replace µ with µ + c and replace each αi with αi − c, the means will not change. When this occurs the parameters are said to be unidentified. To estimate the parameters, we must impose a constraint. In the present situation, the constraint will be
i αi = 0.
This allows the αi to be interpreted as “deviations from the mean” – if α2 = 3, then Atou wheat yields on average three tons more than the overall mean, and if α4 = −2, Mardler wheat yields on average two tons less than the overall mean.
- In order to estimate the population parameters, we use the
same “sum of squared residuals” function that was used for simple linear regression:

SLIDE 7
(Yij − ˆ µ − ˆ αi)2. As with simple linear regression, our estimates will be the val- ues that we get by searching for the values of ˆ µ, ˆ α1, . . . , ˆ α4 that make the sum of squared residuals as small as possi-
- ble. Without derivation, the following are the least squares
parameter estimates: ˆ αi = ¯ Yi· − ¯ Y·· ˆ µ = ¯ Y·· For the example given above we get

SLIDE 8
ˆ α1 = −0.176 ˆ α2 = 0.041 ˆ α3 = 0.129 ˆ α4 = 0.006 ˆ µ = 5.42 Also by analogy with simple linear regression, we can define fitted values ˆ Yij = ˆ µ + ˆ αi residuals

SLIDE 9 rij = Yij − ˆ µ − ˆ αi, and an estimate of the standard deviation ˆ σ =
r2
ij/(mn − m).
- Since EYi1 = · · · = EYin = µ+αi, it follows that E ¯
Yi· = µ+αi. Similarly, E ¯ Y·· = µ+
i αi/m = µ. Thus Eˆ
αi = µ+αi−µ = αi, so ˆ αi is unbiased. Similarly it can be shown that ˆ µ is unbiased.

SLIDE 10
αi can be calculated directly: var(ˆ αi) = var(¯ Yi· − ¯ Y··) = var(¯ Yi·) + var(¯ Y··) − 2cov(¯ Yi·, ¯ Y··) = σ2/n + σ2/mn − 2cov(Yi·, Y··)/n2m = σ2/n + σ2/mn − 2cov(Yi·, Yi·)/n2m = σ2/n + σ2/mn − 2nσ2/n2m = σ2/n + σ2/mn − 2σ2/mn = σ2/n − σ2/mn = σ2 · m − 1 mn .

SLIDE 11
- Based on the variance formula for ˆ
αi, we can carry out hy- pothesis tests. For example, to test α2 = 0 versus α2 > 0, the test statistic would be T = ˆ α2 ˆ σ ·
mn
m − 1. When α2 = 0 (the null hypothesis) T has a tm(n−1) distribu-
- tion. In ANOVA problems it is common for m(n − 1) to be
small, so the normal approximation should not generally be used. In the above example, we get ˆ σ = .71, so the (two sided) test statistics and p-values are as follows:

SLIDE 12 Parameter Estimate |T| p-value α1
0.57 .58 α2 0.041 0.13 .90 α3 0.129 0.42 .68 α4 0.006 0.02 .98 So in the example, none of the coefficients are significantly different from zero – we can not confidently conclude that any variety of wheat is better than the others. As we have seen previously, the test statistic −T can be used to test the alternative hypothesis α2 < 0, and the test statistic |T| can be used to test the alternative hypothesis α2 = 0.
- We can also use the standard deviation formula to get a CI
for any of the αi’s. Since

SLIDE 13 P
α2 − α2 ˆ σ ·
mn
m − 1 ≤ Q(.975)
where Q is the tm(n−1) quantile function, it follows that P
ˆ
α2 − ˆ σQ(.975)
mn ≤ α2 ≤ ˆ α2 + ˆ σQ(.975)
mn
= .95.
The confidence intervals for the example are: α1 (−.84, .48) α2 (−.62, .70) α3 (−.53, .79) α4 (−.65, .67).

SLIDE 14
- As with simple linear regression, we have a “sum of squares
law”: SSTO = SSE + SSR
(Yij − ¯ Y··)2 =
(Yij − ˆ Yij)2 +
(ˆ Yij − ¯ Y··)2, where SSR and SSE are uncorrelated. We can define “mean squares”: Sum of Squares DF Mean square SSTO mn-1 SSTO / (mn-1) SSR m-1 SSR / (m-1) SSE m(n-1) SSE / (m(n-1))

SLIDE 15 Large values of SSR and small values of SSE suggest a good fit to the model. Therefore F = MSR/MSE can be used to test the fit of the model. F has a F distribution with (m − 1, m(n − 1)) degrees of freedom. In the example, the sums of squares are SSTO= 6.2350, SSR= 0.1975, and SSE= 6.0374. The mean squares are MSTO= 0.5196, MSR= 0.0658, and MSE= 0.5031. The F statistic is F = .13079, which gives an insignficant p-value
- f around .94. Thus there is no evidence that any type of
wheat has greater or lesser yield than any other.

SLIDE 16 Unabalanced one-way layout
- The balanced one way layout can easily be generalized to
the unbalanced case, where differing numbers of replicates are made for different factor levels. In this case, we use ni to denote the number of replicates for factor level i, and let N =
i ni denote the total number of
The definitions of Yi· and Y·· are the same as in the balanced case, but now we have ¯ Yi· = Yi·/ni and ¯ Y·· = Y··/N. The definitions of ˆ αi, ˆ µ, ˆ Yij, and rij are the same as in the balanced case.

SLIDE 17 In the unbalanced one-way ANOVA, the αi are identified by requiring
niαi = 0. The standard deviation estimate becomes: ˆ σ =
r2
ij/(N − m).
The variance of ˆ αi becomes: Var(ˆ αi) = σ2(1/ni − 1/N) = σ2N − ni Nni .

SLIDE 18 The test statistic for a hypothesis test αi = 0 versus αi > 0 is T = ˆ αi
N − ni /ˆ σ, which has a tN−m distribution. Since P
αi − αi ˆ σ ·
N − ni ≤ Q(.975)
where Q is the tN−m quantile function, it follows that

SLIDE 19 P
αi − ˆ σQ(.975)
Nni ≤ αi ≤ ˆ αi + ˆ σQ(.975)
Nni
- = .95.
- The sum of squares law is the same as in the balanced case,
however the degrees of freedom must be generalized: Sum of Squares DF Mean square SSTO N-1 SSTO / (N-1) SSR m-1 SSR / (m-1) SSE N-m SSE / (N-m) Note that for everything above, the formulas for the balanced case are special cases of the formulas for the unbalanced case, replacing ni with n, and nm with N.

SLIDE 20
- Here is an example unbalanced one-way layout, along with
its ANOVA table:
Group (X) Response (Y ) Yi· ¯ Yi· 1 3.4 3.7 2.9 3.5 3.2 16.7 3.34 2 4.1 3.8 4.2 12.1 4.03 3 3.5 3.7 3.2 3.9 3.5 3.5 21.3 3.55 4 2.5 3.2 3.1 3.6 3.4 3.1 3.0 3.1 25.0 3.13 5 3.9 3.6 3.8 11.3 3.77 86.4 3.46
Note that n1 = 5, n2 = 3, n3 = 6, n4 = 8, n5 = 3, and N = 25. The standard deviation estimate is ˆ σ ≈ .27. The parameter estimates, t-statistics, p-values, and confi- dence intervals are contained in the following table:

SLIDE 21 Parameter Estimate |T| p-value CI α1
1.06 3.0 × 10−1 (−0.34, 0.11) α2 0.58 3.90 8.9 × 10−4 (0.27, 0.89) α3 0.09 0.97 3.5 × 10−1 (−0.11, 0.30) α4
4.15 4.9 × 10−4 (−0.50, −0.16) α5 0.31 2.10 4.8 × 10−2 (0.01, 0.62) The sum of squares law for the example is SSTO = SSR + SSE 3.78 = 2.29 + 1.50, giving 0.57 as the MSR, 0.07 as the MSE, an F-statistic value
- f F = 7.64, and an F-statistic p-value of ≈ 6.5 × 10−4.

SLIDE 22 Since the F-statistic is highly significant, we can reject the null hypothesis that α1 = α2 = α3 = α4. Thus some of the factor levels have different mean values. Based on the hypothesis tests for each αi, we are confident that α2 > 0 and α4 < 0.
Balanced two-way layout
- Suppose now that there are two qualitative X variables (two
factors). Thus each observation has the form (X1, X2, Y ). Let m1 be the number of levels of the first factor, and let m2 be the number of levels of the second factor. Thus there are m1m2 distinct combinations of factor levels. Also suppose that there is exactly one observation made at each combination of levels.

SLIDE 23
For example, in the following data m1 = 4 and m2 = 5. One way to display the data is in the following table: X1 X2 Y X1 X2 Y 1 1 4 3 1 5 1 2 5 3 2 4 1 3 6 3 3 7 1 4 5 3 4 5 1 5 4 3 5 4 2 1 3 4 1 4 2 2 2 4 2 4 2 3 4 4 3 6 2 4 3 4 4 4 2 5 2 4 5 4

SLIDE 24
For example, X1 may indicate 4 different levels of fertilizer (e.g. no, low, medium, high), X2 may indicate 5 different varieties of wheat, and Y may indicate the yield. A more informative way to display this data is as follows: 1 2 3 4 5 1 4 5 6 5 4 2 3 2 4 3 2 3 5 4 7 5 4 4 4 4 6 4 4 X1 X2

SLIDE 25
We may refer to the Y values using the X1 and X2 values as subscripts – for instance when X1 = 3 and X2 = 4, we have Y34 = 5, or when X1 = 2 and X2 = 1, we have Y21 = 3. We will need the row sums Yi· =
j Yij and the column sums
Y·j =
i Yij.
Similarly we will need the row means ¯ Yi· = Yi·/m2 and the column means ¯ Y·j = Y·j/m1. Finally we will need the overall sum Y·· =
ij Yij, and the overall mean
¯ Y·· = Y··/m1m2. All of these quantities can be displayed in the following ANOVA table:

SLIDE 26 4 5 6 5 4 24 4.8 3 2 4 3 2 14 2.8 5 4 7 5 4 25 5.0 4 4 6 4 4 22 4.4 16 15 23 17 14 85 4.00 3.75 5.75 4.25 3.50 4.25 For example, Y2· = 14, ¯ Y4· = 4.4, Y·2 = 15, ¯ Y·4 = 4.25, Y·· = 85, and ¯ Y·· = 4.25.
- We will consider several different models for this data. The
central model is the additive model, that specifies the fol- lowing mean values for each observation: EYij = µ + αi + βj.

SLIDE 27 The unknown parameters are α1, . . . , αm1, β1, . . . , βm2, µ, and σ. Looking at another small example, suppose that m1 = 2, m2 = 3, the population values are µ = −1, α1 = 3, α2 = −3, β1 = −5, β2 = 0, β3 = 5. The interior of the following “table of means” shows the mean values for each Yij, and the margins show the population αi and βj values.
5 3
2 7
1 The additive model is

SLIDE 28 Yij = µ + αi + βj + ǫij where the ǫij are iid random variables with mean 0 and stan- dard deviation σ. The following table shows each observed value expressed as its mean value plus an error term:
5 3 −3 + ǫ11 2 + ǫ12 7 + ǫ13
−9 + ǫ21 −4 + ǫ22 1 + ǫ23
- In practice we won’t know the αi, βj, µ, and σ values, so we
will estimate them by minimizing the least squares function:

SLIDE 29
(Yij − ˆ µ − ˆ αi − ˆ βj)2. As with the one-way layout, we are not able to identify the parameters – adding a constant to every αi and subtracting that constant from every βj does not change the mean levels. To be able to identify the parameters, we require
αi = 0 and

SLIDE 30
βj = 0. The least squares parameter estimates are: ˆ µ = ¯ Y·· ˆ αi = ¯ Yi· − ¯ Y·· ˆ βj = ¯ Y·j − ¯ Y·· For the 4 × 5 example above, the parameter estimates and fitted values are given in the following “table of fitted val- ues”:

SLIDE 31 4.55 4.3 6.3 4.8 4.05 0.55 2.55 2.3 4.3 2.8 2.05
4.75 4.5 6.5 5.0 4.25 0.75 4.15 3.9 5.9 4.4 3.65 0.15
1.5
4.25 The residuals are given in the following “table of residuals”:
0.70
0.20
0.45
0.20
0.25
0.50 0.00
0.10 0.10
0.35 As with any least squares fit, you can check that the residuals sum to zero.

SLIDE 32
- Applying a similar derivation as was used in the one-way
layout, the variances of the parameter estimates are: var(ˆ αi) = σ2(1/m2 − 1/m1m2) = σ2m1 − 1 m1m2 . var(ˆ βj) = σ2(1/m1 − 1/m1m2) = σ2m2 − 1 m1m2 . It can also be easily shown that these estimates are unbiased: so Eˆ αi = αi and E ˆ βj = βj. We will also need to study the correlation between ˆ αi and ˆ βj, for any pair of indices i, j:

SLIDE 33 cov(ˆ αi, ˆ βj) = cov(¯ Yi· − ¯ Y··, ¯ Y·j − ¯ Y··) = σ2/m1m2 − σ2/m1m2 − σ2/m1m2 + σ2/m1m2 = 0. Thus ˆ αi and ˆ βj are uncorrelated.
ˆ Yij = ˆ µ + ˆ αi + ˆ βj = ¯ Yi· + ¯ Y·j − ¯ Y··, the residuals are

SLIDE 34 rij = Yij − ¯ Yi· − ¯ Y·j + ¯ Y··, and the error standard deviation estimate is ˆ σ =
r2
ij/(m1 − 1)(m2 − 1).
- The usual “sum of squares” law applies:
- ij
(Yij − ¯ Y··)2 =
(Yij − ˆ Yij)2 +
(ˆ Yij − ¯ Y··)2, but now we can take it a step further, decomposing SSR as:

SLIDE 35
(ˆ Yij − ¯ Y··)2 =
(ˆ αi + ˆ βj)2 = m2
ˆ α2
i + m1
ˆ β2
j +
ˆ αiˆ βj. Since
i ˆ
αi =
j ˆ
βj = 0, it follows that
ˆ αiˆ βj =
ˆ αi
ˆ βj = 0. Thus we can decompose SSR = SSA + SSB, where SSA = m2
ˆ α2
i

SLIDE 36 and SSB = m1
ˆ β2
j .
Thus for the balanced, two-way layout we get: SSTO = SSE + SSA + SSB. This decomposition can be used to carry out several tests regarding the structure of the model. Before we see how to do this, the degrees of freedom and mean squares are as follows (where N = m1m2 is the total sample size):

SLIDE 37
SS MS DF SSTO MSTO N − 1 SSR MSR m1 + m2 − 2 SSA MSA m1 − 1 SSB MSB m2 − 1 SSE MSE (m1 − 1)(m2 − 1) For the example given above, the sums of squares and mean squares are: SSTO 29.75 MSTO 1.57 SSR 27.45 MSR 3.92 SSA 14.95 MSA 4.98 SSB 12.50 MSB 3.13 SSE 2.30 MSE 0.19

SLIDE 38
- We can use the sum of squares to assess the model that best
fits the data. First note that every two-way layout contains two one-way layouts as special cases. If α1 = · · · = αm1 = 0, then the model reduces to the one-way layout EYij = µ + βj (the i index doesn’t affect the mean). Similarly, if β1 = · · · = βm2 = 0, then the model reduces to EYij = µ + αi (the j index doesn’t affect the mean). If SSA is small, then it may easily be true that all αi = 0, while if SSA is large, the data strongly suggest that some
Thus we can use SSA as our test statistic for the null hypothesis α1 = · · · = αm1 = 0. We normalize to get an F-statistic: F = MSA MSE,

SLIDE 39
with m1 − 1, (m1 − 1)(m2 − 1) DF. In the example, we get F ≈ 26.21, giving a p-value < 10−5. Similarly, to test the null hypothesis β1 = · · · = βm2 = 0, use F = MSB MSE, which has m2 − 1, (m1 − 1)(m2 − 1) DF. In the example, we get F ≈ 16.47, giving a p-value < 10−4. We may also test the null hypothesis that all αi = 0 and all βj = 0 using F = MSR MSE,

SLIDE 40 which has m1 + m2 − 2, (m1 − 1)(m2 − 1) DF. In the example, we get F ≈ 20.63, giving a p-value < 10−5.
Balanced two-way layout with replicates
- Suppose, as above, that there are two factors influencing
the response, so the data takes the form (X1, X2, Y ). Now suppose that for each combination of X1&X2, we observe r > 1 replicates of Y .

SLIDE 41
For example, the following data come from an experiment where the response (Y ) is the weight of 16 week old chicks, and the factors are “level of protein” (X1) and “level of fish solubles” (X2). There are m1 = 3 levels of protein (low, medium, high), and m2 = 2 levels of fish solubles (absent, present). For each combination of factor levels, there are r = 2 replicates. The data can be written: No FS FS Low 7094, 7053 8005, 7657 Med 6943, 6249 7359, 7292 High 6748, 6422 6764, 6560 We use Yijk to denote replicate k within cell i, j (i.e. replicate k where X1 = i and X2 = j). For example, Y111 = 7094, and Y112 = 7053.

SLIDE 42
- Row and column sums, Yi··, Y·j·, and the total Y··· now include
a sum over the replicates as well (hence the third · in the subscript). The row and column means, and grand mean must account for the replicates in their denominator, so ¯ Yi·· = Yi··/m2r, ¯ Y·j· = Y·j·/m1r, and ¯ Y··· = Y···/m1m2r. The following table shows the data given above, with the row and column sums and averages given in the margins. No FS FS Low 7094, 7053 8005, 7657 29809 7452 Med 6943, 6249 7359, 7292 27843 6961 High 6748, 6422 6764, 6560 26494 6624 40509 43637 84146 6752 7273 7012

SLIDE 43 There is also a new average, the “within cell average” ¯ Yij· formed by averaging all values in a single cell. The table of within cell averages for the example follows: No FS FS Low 7074 7831 Med 6596 7326 High 6585 6662
- Parameter estimates are defined as in the case with no repli-
cates, but the standard errors will be smaller since we have more data when r > 1:

SLIDE 44 Parameter Estimate Variance αi ¯ Yi·· − ¯ Y··· σ2(m1 − 1)/N βj ¯ Y·j· − ¯ Y··· σ2(m2 − 1)/N µ ¯ Y··· σ2/N Fitted values are defined as ˆ Yijk = ˆ µ + ˆ αi + ˆ βj. Note that the fitted values for all replicates in a common cell are identical (i.e. ˆ Yijk does not depend on k).
- By direct analogy with the two-way layout with no replicates,
we have the sum of squares law SSTO = SSE + SSA + SSB (note that SSA and SSB are scaled by r in this case):

SLIDE 45
Y )2 =
Yijk)2 + m2r
ˆ α2
i + m1r
ˆ β2
j .
Since we have replicates, we can further decompose SSE:
Yijk)2 =
Yij· + ¯ Yij· − ˆ Yijk)2 =
Yij·)2 +
Yij· − ˆ Yijk)2 +2
Yij·)(¯ Yij· − ˆ Yijk). Focusing on the third term, and using the fact that ˆ Yijk does not depend on k,

SLIDE 46
(Yijk − ¯ Yij·)(¯ Yij· − ˆ Yijk) =
(¯ Yij· − ˆ Yijk)
(Yijk − ¯ Yij·) = Thus we have a decomposition SSE = SSP + SSI, where SSP =
Yij·)2 SSI =
Yij· − ˆ Yijk)2,

SLIDE 47 where SSP is the “pure error” sum of squares, and SSI is the “interaction” sum of squares. Note that when r = 1, SSP = 0 and SSI = SSE.
- Degrees of freedom and mean squares are as follows:
Sum Mean DF SSTO MSTO N − 1 SSR MSR m1 + m2 − 2 SSA MSA m1 − 1 SSB MSB m2 − 1 SSE MSE N − m1 − m2 + 1 SSP MSP m1m2(r − 1) SSI MSI (m1 − 1)(m2 − 1)

SLIDE 48
Note that when r = 1, SSP has 0 DF, and the DF’s for SSI and SSE are identical. For the example, the sums of squares and degrees of freedom are: Sum DF SSTO 2879821 11 SSR 2204881 3 SSA 1389515 2 SSB 815365 1 SSE 674941 8 SSP 378401 6 SSI 296540 2 The usual estimate for the error standard deviation is

SLIDE 49 ˆ σ =
ijk/(N − m1 − m2 + 1),
but we also have a different estimate ˆ σpure =
Yij· − Yijk)2/m1m2(r − 1). In the example, ˆ σ = 290 while ˆ σpure = 251.
- As in the case r = 1, we have F-tests for the null hypothesis
“all αi = 0” (F = MSA / MSE), for the null hypothesis “all βj = 0” (F = MSB / MSE), and for the null hypothesis “all αi = 0 and all βj = 0” (F = MSR / MSE).

SLIDE 50 In the example, MSA/MSE ≈ 8.2, giving a p-value ≈ .01, and MSB/MSE ≈ 9.7, also giving a p-value ≈ .01. So both the row and column effects are significantly different from zero.
- When r > 1 we are also able to test the additivity hypothesis
(i.e. that EYijk = µ + αi + βj, as opposed to EYijk = θij, where the θij are completely unconstrained). For example, the table of population means on the left is additive, while that on the right is not: 3 5 1 2 4 4 3 2 3 4

SLIDE 51 Before we consider formal tests, it is useful to construct some simple plots that suggest whether the effects are interactive. The following are plots of the points (j, ¯ Yij·), where points with a common i value are connected by lines. The plot on the left corresponds to the table of means on the left, above, and the plot on the right corresponds to the table of means
1 2 3 4 5 0.5 1 1.5 2 0.5 1 1.5 2 2.5 3 3.5 4 0.5 1 1.5 2

SLIDE 52 If the the lines are approximately parallel, then additivity is
- suggested. In the above plots, the left side is perfectly addi-
tive, while the right side is strongly non-additive. Similarly, we can plot (i, ¯ Yij·), connecting points with a common j value.
2 2.5 3 3.5 4 4.5 5 0.2 0.4 0.6 0.8 1 0.5 1 1.5 2 2.5 3 3.5 4 0.2 0.4 0.6 0.8 1
The same conclusion is suggested.

SLIDE 53 Here are the two diagnostic plots for the chick weight exam- ple data:
6400 6600 6800 7000 7200 7400 7600 7800 8000 0.2 0.4 0.6 0.8 1 6400 6600 6800 7000 7200 7400 7600 7800 8000 0.5 1 1.5 2
There is some evidence of non-additivity, but it is not ex- tremely strong.

SLIDE 54
- Using sample means rather than population means, additivity
will not hold exactly due to random variation, so we need to assess whether the means are sufficiently close to additive to infer that the underlying population means are exactly addi-
- tive. This is acccomplished using the F-test F = MSI/MSP.
In the chick weight example, the F-statistic is ≈ 2.4, giving a p-value of ≈ .17. This provides quantitative evidence that the interactivity suggested in the plot above is not unexpected given the sample size and noise level.