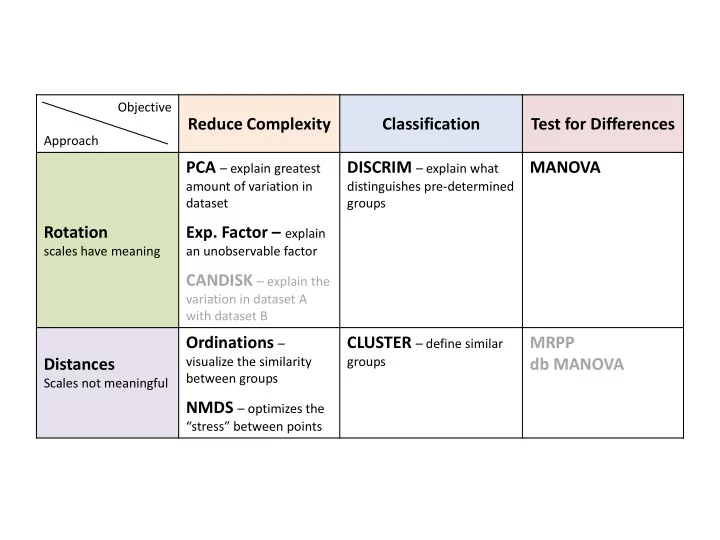
Objective Approach
Reduce Complexity Classification Test for Differences Rotation
scales have meaning
PCA – explain greatest
amount of variation in dataset
- Exp. Factor – explain
an unobservable factor
CANDISK – explain the
variation in dataset A with dataset B
DISCRIM – explain what
distinguishes pre-determined groups
MANOVA Distances
Scales not meaningful
Ordinations –
visualize the similarity between groups
NMDS – optimizes the
“stress” between points
CLUSTER – define similar
groups
MRPP db MANOVA