Methods for calculating rare event dynamics and pathways of - - PowerPoint PPT Presentation
Methods for calculating rare event dynamics and pathways of - - PowerPoint PPT Presentation
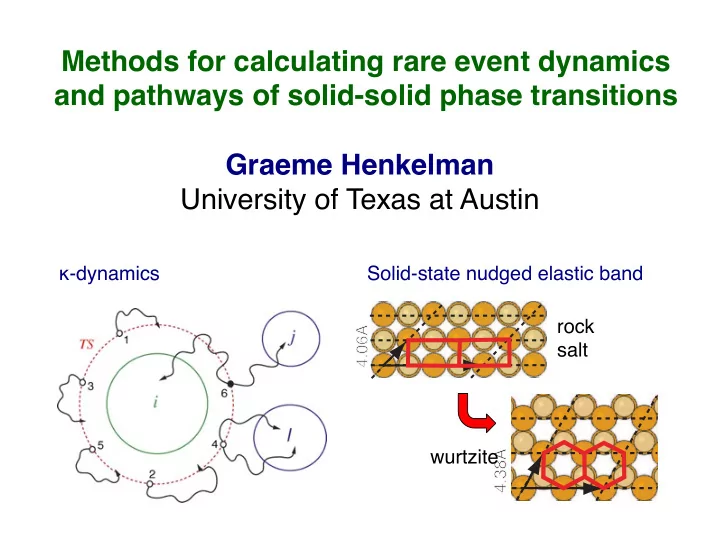
Methods for calculating rare event dynamics and pathways of solid-solid phase transitions Graeme Henkelman University of Texas at Austin -dynamics Solid-state nudged elastic band rock 4.06 4.06 salt wurtzite 4.38 4.38 Classes
Classes of Dynamical Systems
Rough Smooth Energy Landscape: Final State: Unknown Known
Initial Final Initial Initial Final
?
Transition state theory
A statistical theory for calculating the rate of slow thermal processes -- rare event dynamics Requires an N-1 dimensional dividing surface that is a bottleneck for the transition:
Harmonic transition state theory
Find saddle points on the energy surface Rate of escape through each saddle point region:
kHTST = QN
i=1 νi
QN−1
j=1 ν† j
exp ✓ − ∆E kBT ◆
∆E
R P
x = x†
kTST = 1 2 ⌦ δ(x − x†)|v⊥| ↵
R
KMC: the Good, the Bad, and the Ugly
The Good For rare event systems where transition rates are defined by first-order rate constants, KMC is an exact stochastic solution to the kinetic master equation. The Bad It is very hard to determine all possible kinetic events available to the simulation. It is very hard to calculate an exact rate for a process, let alone all of them. Typically limited to transition state theory (e.g. harmonic TST within aKMC). The Ugly It’s a lot of work calculating all possible events and rates to find one trajectory. where where μ is random on (0,1]
Dynamical corrections to TST
TST assumes that all trajectories that cross the TS are reactive trajectories. : the ratio of successful trajectories to number of crossing points ratio between the TST and true rate: both total escape rate and by product: A successful trajectory: 1) trajectory must go directly to products without recrossing the TS 2) trajectory must start in initial state Example: violation
- f TST
Dynamical correction factor
The κ-dynamics algorithm
- 1. Choose a reaction coordinate and
sample a transition state (TS) surface
- 2. Launch short time trajectories until
- ne goes directly to a product and starts
in the initial state; record the number, N
- 3. If no successful trajectory within Nmax,
push TS up in free energy and go to 2
- 4. Calculate kTST for successful TS
surface (parallel tempering, WHAM)
- 5. Update the simulation clock by
where μn are random numbers on (0,1]
- 6. Repeat procedure in product state
1 2 3 4 5 6 7
C.-Y. Lu, D. E. Makarov, and G. Henkelman,
- J. Chem. Phys. 133, 201101 (2010).
Sketch of proof that κ-dynamics is exact
(1) Branching ratio The probability of reaching state j is:
- A. Voter and J. D. Doll JCP 82, 1 (1985)
where sampled (2) Reaction time where is the true rate What we want: What we have in κ-dynamics: where (1 success)(N-1 failures) Connection with TST rate: Erlang N-distribution Combining: the correct distribution based upon the true rate
Reaction coordinate test
TST+κ hTST
Al/Al(100), hop on frozen surface
kTST
Exchange on relaxed surface
κ-dyn
κ-dynamics rates are correct and independent of reaction coordinate
κ-dynamics
Bond-boost: specifies the stretch in the most-stretched bond:
- R. A. Miron and K. A. Fichthorn, JCP 119, 6210 (2003)
Bond stretching parameter
Branching ratio test
Comparison of reaction products for classical dynamics and κ-dynamics Al/Al(100), relaxed surface
Reaction mechanism Branching ratio direct MD κ-dynamics harmonic TST 0.0 0.5 1.0 1 2 3 4 5 6 7 8 1 2 3 4 5 6 7 8
- 200K
400K
κ-dynamics trajectories
Al/Al(100) island ripening Al/Al(100) pyramid collapse
Solid-solid phase transitions
4.06Å 4.06Å
4.38Å 4.38Å
rock salt wurtzite Some transitions involve both change in atomic and cell degrees of freedom E.g. CdSe: zite
Potential parameters: E. Rabani, J. Chem. Phys. 116, 258 (2002).
Nudged Elastic Band Method
zite Pratt, Elber, Karplus, ... and others
Initial Final
F τ F FS F
NEB
MEP NEB
i i i i i i i+1 i−1
ˆ
Pioneering work Nudged elastic band
[1] H. Jónsson, G. Mills, and K.W. Jacobsen, in Classical and Quantum Dynamics in Condensed Phase Simulations, 385 (1998). [2] G. Henkelman and H. Jónsson, J. Chem. Phys. 113, 9978 (2000). [3] G. Henkelman, B. P. Uberuaga, and H. Jónsson, J. Chem. Phys. 113, 9901 (2000).
Images connect initial and final states Perpendicular component (potential): NEB force on each image: Parallel component (springs):
Danger of assuming a reaction coordinate
final initial energy reaction coordinate
drag NEB
x 1
- 1
- 2
2 3 1 rAB saddle initial final i n i t i a l b a n d NEB drag drag direction
The x-coordinate separates initial from final state, but is not a suitable reaction coordinate: Initial Final A drag calculation along x (black points) misses the saddle point where the reaction coordinate follows the y-direction x y reaction coordinate The resulting barrier is under-estimated:
Cell variables vs atomic coordinates
x 1
- 1
- 2
2 3 1 rAB saddle initial final initial band NEB drag drag direction
a) atom dominated b) cell dominated final (wurtzite)
4.38 Å 4.22 Å 4.06 Å 8.75 Å 9.36 Å 9.08 Å 10.98 Å 11.49 Å
initial (rock salt) transition state
8.75 Å 4.20Å 4.20Å 4.20Å 4.06Å 4.06Å 4.06Å 4.38Å 4.38Å 4.38Å
atom dominated cell dominated
Two pathways for the same solid-solid phase transition in CdSe
cell
NEB calculations in only atomic [1] or cell (RNM) [2] coordinates have the errors found in drag
- methods. Requires a
unified approach
atoms
[1] D. R. Trinkle, R. G. Hennig, S. G. Srinivasan, et al., PRL 91, 025701 (2003). [2] K. J. Caspersen and E. A. Carter, PNAS 102, 6738 (2005).
Choice of cell representation
Cell coordinates Choose a representation which does not allow for net rotation of the lattice Changes in the cell are represented as strain
v3 = (h ,h ,h )
3x 3y 3z
v1 = (h ,0,0)
1x
= (h ,h ,0)
2x 2y
v2
periodic solid expanded periodic solid ɛ = δ 0
( )
0 δ
Strain describes changes in the lattice which is invariant to the unit cell
Combining atomic and cell variables (G-SSNEB)
Single displacement vector: Jacobian to combine different units change in atom positions change in cell shape Requirement: reaction pathways should be independent of unit cell size and shape -- the path should be a property of the infinite solid Unit of length, average distance between atoms: Scaling of atomic displacements with size: Choose J so that Jε has same units and scaling as ΔR: Similar logic applies to the stress / force vector: volume
Results
NEB
15 meV
energy / atom (meV)
- 60
- 40
- 20
20
13 meV
initial final initial state final state saddle G-SSNEB RNM NEB G-SSNEB RNM saddle
12 meV 16 meV
energy / atom (meV) initial final NEB saddle G-SSNEB RNM initial state final state G-SSNEB NEB RNM
- 60
- 40
- 20
20 saddle
Atom-dominated process: Cell-dominated process: RNM fails; NEB / G-SSNEB work NEB fails; RNM / G-SSNEB work
Scaling with system size
energy / atom (meV)
- 60
- 40
- 20
20
- 1
1
- 2
distance / N (Å) 8 32 72 128 atoms / cell 128 transition state
Minimum energy path is insensitive to unit cell size and shape:
Change of mechanism
With increasing system size:
b
a
energy (eV) number of atoms in unit cell 10 10,000 1,000 100 0.1 1 10 c b a
The mechanism changes from a concerted bulk process (cell dominated) to a local (atom dominated) 3d (bulk concerted) 2d (line defect) 1d (point defect) a
Movies
Concerted mechanism Local mechanism
Local process in detail
- 80
- 60
- 40
- 20
energy / atom (meV) 1 2 initial final a c b
c b a
Complex mechanism:
Research Group
Chun-Yaung (Albert) Lu Daniel Sheppard Penghao Xiao
Software tools Research Group and Collaborators
Chun Yaung Lu (graduate student, now at LANL) Daniel Sheppard (graduate student, now at LANL) Penghao Xiao (graduate student) Rye Terrell (graduate student) Sam Chill (graduate student) Will Chemelewski (graduate student) Dmitrii Makarov (U. Texas, Austin) Hannes Jónsson (U. Iceland) Andreas Pederson (U. Iceland) Duane Johnson (Iowa State/Ames Lab) DOE - EFRC, SISGR NSF - CAREER, NIRT Welch Foundation AFOSR
Funding
Acknowledgments
http://theory.cm.utexas.edu/vtsttools/ AKMC, Dimer, (SS)NEB, and dynamical matrix
- methods implemented in the VASP code