An in-house expression database : CleanEx CleanEx : CONCEPT AND - PowerPoint PPT Presentation
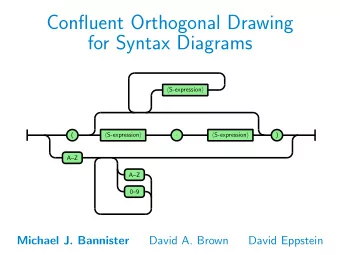
An in-house expression database : CleanEx CleanEx : CONCEPT AND ORGANIZATION CleanEx_exp CleanEx_trg CleanEx BUILDING CleanEx Material : source databases CleanEx_exp files CleanEx_trg files CleanEx link file CleanEx : Main objectives
An in-house expression database : CleanEx CleanEx : CONCEPT AND ORGANIZATION CleanEx_exp CleanEx_trg CleanEx BUILDING CleanEx Material : source databases CleanEx_exp files CleanEx_trg files CleanEx link file
CleanEx : Main objectives & data organization To give access to heterogeneous expression data concerning the same gene through the same name --> The CleanEx file type To reformat these heterogenous data in a way that will allow joint analysis and cross-dataset comparisons --> The CleanEx_exp file type To allocate expression results of unknown sequences to the corresponding approved gene name once it is known -- > The CleanEx_trg file type To provide a weekly updated annotation of so-called “targets” via an adapted mapping procedure
What should be in there ? Mandatory information Experiment meta-data (clinical information, scanner settings, tools used for normalization, protocol, organism, sample preparation...) Chip meta-data : spot-to-gene, or at least spot-to-sequence information Gene expression numerical data (for each feature and for each sample) In-house specific identifiers for data retrieval ( Samples, Chips, Datasets) Optional information Link to other databases List of medical keywords for each experiment Associated datasets Reformatted numerical data (log2, re-normalized...)
Source databases to build CleanEx The construction procedure is based on the official organism’s gene catalog and it’s corresponding UniGene clusters. Gene nomenclature official lists : HUGO nomenclature in the Genew database for human MGD nomenclature for Mouse Unigene : clusters of transcript sequences coming from the same locus. mRNA sequences databases : RefSeq : set of high-quality curated mRNA sequences mRNAs from GenBank HTCs from GenBank ESTs Gene Expression Omnibus : expression data in “soft” format
Structure of the CleanEx database Data are stored in three different file formats : 1- CleanEx_exp, the reformatted expression data file. 2- CleanEx_trg : contains the mapping ot the « expression targets » to the approved genes symbols. 3- CleanEx, linked to CleanEx_exp and CleanEx_trg via clones AC, RNAs or RefSeq ACs and cross-referenced with external databases.
Structure of the CleanEx database Praz et al. Nucleic Acids Res. 32:D542-D547(2004)
CleanEx_exp : structure Contains the downloaded expression data. Heterogeneous public data are downloaded and first submitted to a quality control. Each dataset is reformatted in a way to preserve all relevant information from the original sources and according to the data type. Each dataset produces a « meta-entry » in the CleanEx_exp file type. Each entry stores the measurements of one « expression target » for all the experiments done in the dataset.
CleanEx_exp : formatting procedure One Experiment, all targets One target, all experiments Target 1 Experiment 1 Exp_1 Result_1 Exp_1 Trg_1 Result_1 Exp_1 Exp_2 Result_1 Exp_2 Trg_2 Result_2 Exp_1 Exp_3 Result_1 Exp_3 Trg_3 Result_3 Exp_1 Target 2 Experiment 2 Exp_1 Result_2 Exp_1 Trg_1 Result_1 Exp_2 Exp_2 Result_2 Exp_2 Trg_2 Result_2 Exp_2 Exp_3 Result_2 Exp_3 Trg_3 Result_3 Exp_2 Target 3 Experiment 3 Result_3 Exp_1 Exp_1 Trg_1 Result_1 Exp_3 Exp_2 Result_3 Exp_2 Trg_2 Result_2 Exp_3 Exp_3 Result_3 Exp_3 Trg_3 Result_3 Exp_3
CleanEx_exp : dual channel experiments integration
CleanEx_ep : Affymetrix experiments integration
CleanEx_trg : content and build Contains the link between “targets” submitted to experiments stored in CleanEx_exp and the existing approved gene symbols. Provides a « quality criteria » to assess the reliability of the target (clone, tag, probeset...) regarding it’s corresponding gene. Is updated each time the gene catalog is changed. The update procedure depends on the target type.
Raw data generation details : Affymetrix From : http://www.affymetrix.com
Raw data generation details : SAGE and MPSS From : http://www.ncbi.nlm.nih.gov/Class /NAWBIS/Modules/Expression From : http://www.lynxgen.com
CleanEx_trg : update procedure For clones : direct mapping to UniGene clusters via EMBL accession numbers. For Affymetrix probesets, SAGE tags, oligos..., we use a two-steps procedure which includes a re-mapping of the tags’ sequences on the RefSeq database. Gene symbol U Clone, EST... n UG_ID i Description g RNA_AC e Clone_AC n RefSeq TAGGER RefSeq/mRNA e GeneID Affy, SAGE...
The tagger program - Designed to search for matches between large collections of short (14–30 nucleotides) words and full genomes or transcriptomes sequence databases. Generates a table index of 13 nucleotides long words and then searches for matches in the sequence database - --> Optimal solution for finding exact matches of Affymetrix probes, MPSS or SAGE tags The tagger and the fetchGWI tools are available online at : http://www.isrec.isb-sib.ch/tagger/
CleanEx_trg : Affymetrix update procedure
CleanEx_trg : SAGE and MPSS update
CleanEx_trg : quality tag The 4 quality levels in CleanEx for Affy, SAGE and MPSS High : All the features of the target correspond to a maximum of two gene clusters. Medium : All the features of the target correspond to a maximum of four gene clusters. Three mismatches are allowed. Low : Criteria are below the ones of the "Medium" tag. Unknown : The target does not yet belong to a Unigene cluster.
CleanEx : the link file Cleanex is a gene index with hyperlinks to external databases and cross-references to expression data in CleanEx_ref. It contains one entry per officially approved gene. It is based on an authoritative reference gene catalogue for each organism considered. For human we use Genew, the gene nomenclature database of HUGO. For mouse, we use the MGD database • It is updated each time CleanEx_trg is changed (weekly).
Expression data External public databases h ftp ttp Data repository h ftp ftp ftp ttp reforma t CleanEx_trg HUGO Unigene Swissprot EPD CleanEx CleanEx_ref Gene Gene Gene Gene Gene Gene symbol symbol symbol symbol symbol symbol UG_ID UG_ID Descripti Descripti on on RNA_AC RNA_AC Clone_AC Clone_AC Refseq_A Refseq_A C C LocusLin LocusLin k k SP_ID+A SP_ID+A C C EPD_ID EPD_ID Exp data CleanEx : updating procedures Exp_ID Exp_ID Exp_ID Target_ID Target_I D
CleanEx : web-based interfaces Single entry search engines CleanEx viewer CleanEx_Exp : expression viewer CleanEx_trg Batch search for CleanEx_trg Cross dataset analysis Step-by-step expression pattern search Common genes retrieval Retrieving expression data Data extraction from one dataset Data from different datasets Using the MeSH terms to extract specific data
Using CleanEx : single entry retrieval Sequence Clones GENE ENTRY mRNAs External Links Expression data
Using CleanEx : single entry retrieval
Using CleanEx : single entry retrieval
Using CleanEx : single entry retrieval
Using CleanEx : Target retrieval
Using CleanEx : Target retrieval
Using CleanEx : Target batch search
Using CleanEx : Target batch search
Using CleanEx : MeSH terms index Key question : how to retrieve biological- and medical-specific expression data ? Medical Subject Headings (MeSH) - a controlled vocabulary by the National Library of Medicine used for indexing and searching for biomedical and health-related information. -Terms are arranged in a hierarchical (tree) structure. -Each expression dataset in CleanEx has been annotated using the MeSH terms list --> rapid access to expression data having a certain biological or medical specificity
Using CleanEx : extracting data - Direct access to a list of datasets related to specific keywords (MeSH or general search) - Specific dataset access by “walking down” the MeSH terms tree - Experiment selection and filters - Generation of two data pools for further comparison - Finding Common Genes List across datasets
Using CleanEx : step-by-step analysis Example : comparing gene expression levels in low grade versus high grade astrocytomas Expression dataset 1 High-grade Low-grade VS Over-expressed genes Extract gene list View genes Retrieve sequences Continue analysis ISREC Ontologizer SSA CleanEx step 1 CleanEx step 2
Using CleanEx : step-by-step analysis
Using CleanEx : step-by-step analysis
Recommend
More recommend
Explore More Topics
Stay informed with curated content and fresh updates.