23/02/16 1
WormBase ParaSite Workshop
Glasgow 24th February 2016
WormBase ParaSite Team
Bruce Bolt BioinformaCcian (web and tools) Jane Lomax BioinformaCcian (curaCon) Myriam Shafie BioinformaCcian (pipelines) Kevin Howe WormBase Team Leader Paul Kersey PI (at EMBL-EBI) Ma< Berriman PI (at Sanger InsCtute)
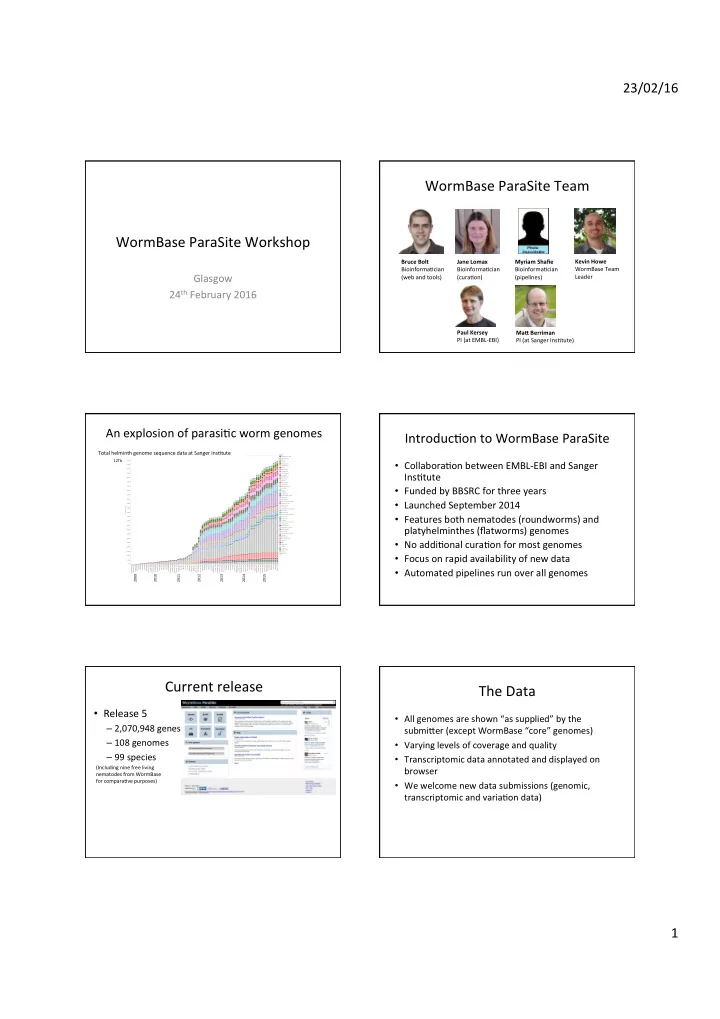
An explosion of parasiCc worm genomes
Run date September 2008 December 2008 January 2009 March 2009 April 2009 May 2009 June 2009 July 2009 August 2009 September 2009 October 2009 November 2009 December 2009 January 2010 February 2010 March 2010 April 2010 May 2010 June 2010 July 2010 August 2010 September 2010 October 2010 November 2010 December 2010 February 2011 March 2011 April 2011 May 2011 June 2011 July 2011 August 2011 September 2011 October 2011 November 2011 December 2011 January 2012 February 2012 March 2012 April 2012 May 2012 June 2012 July 2012 August 2012 September 2012 October 2012 November 2012 December 2012 January 2013 February 2013 March 2013 April 2013 May 2013 June 2013 July 2013 August 2013 September 2013 October 2013 November 2013 December 2013 January 2014 February 2014 March 2014 April 2014 May 2014 June 2014 August 2014 September 201.. October 2014 November 2014 December 2014 January 2015 February 2015 March 2015 April 2015 May 2015 June 2015 July 2015 August 2015 0B 500B 1000B 1500B 2000B 2500B 3000B 3500B 4000B 4500B 5000B 5500B 6000B 6500B 7000B 7500B 8000B 8500B 9000B 9500B 10000B 10500B 11000B 11500B 12000B Cumulative bases Cumulative bases sequenced for helminth tracking Genus Allobilharzia visceralis Angiostrongylus Anisakis Ascaris Atractolytocestus Austrobilharzia Brugia Caenorhabditis Cyathostomum Cylicostephanus Dicrocoelium Diphyllobothrium Dracunculus Dugesia Echinococcus Echinostoma Elaeophora elaphi Enterobius Fasciola Globodera Gongylonema Griphobilharzia amoena Haemonchus Halicephalobus Haplobothrium globuliforme Heligmosomoides Homo sapiens Hymenolepis Macrobilharzia macrobilharzia Mansonella Mesocestoides Nippostrongylus brasiliensis Onchocerca Opisthorchis Parascaris Parastrongyloides trichosuri Protopolystoma Rhabditophanes Sanguinicola cf. inermis Schistocephalus Schistosoma Schistosomatium douthitti Spirometra Strongyloides Strongylus vulgaris Syphacia Taenia Teladorsagia Thelazia Toxocara Trichobilharzia Trichuris WuchereriaTotal helminth genome sequence data at Sanger InsCtute
2009 2010 2011 2012 2013 2014 2015 12Tb
IntroducCon to WormBase ParaSite
- CollaboraCon between EMBL-EBI and Sanger
InsCtute
- Funded by BBSRC for three years
- Launched September 2014
- Features both nematodes (roundworms) and
platyhelminthes (flatworms) genomes
- No addiConal curaCon for most genomes
- Focus on rapid availability of new data
- Automated pipelines run over all genomes
Current release
- Release 5
– 2,070,948 genes – 108 genomes – 99 species
(Including nine free living nematodes from WormBase for comparaCve purposes)
The Data
- All genomes are shown “as supplied” by the
submi]er (except WormBase “core” genomes)
- Varying levels of coverage and quality
- Transcriptomic data annotated and displayed on
browser
- We welcome new data submissions (genomic,