SLIDE 1
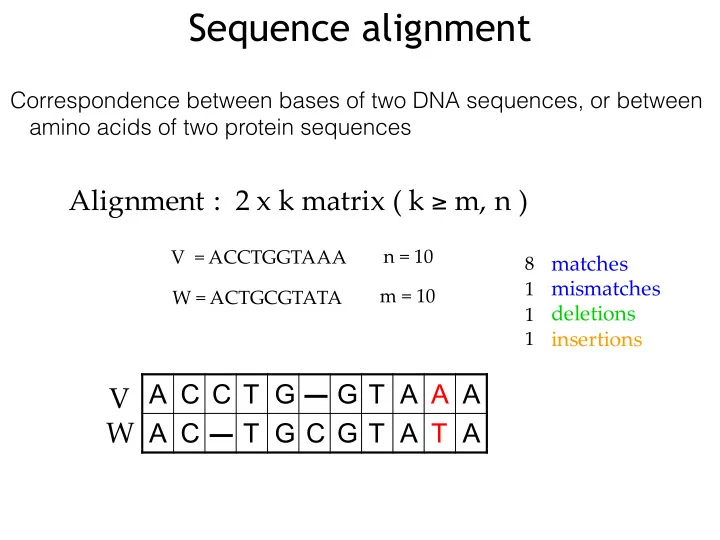
Correspondence between bases of two DNA sequences, or between amino acids of two protein sequences
Sequence alignment
V""="ACCTGGTAAA W"="ACTGCGTATA n"="10 m"="10
A C C T G G T A A A A C T G C G T A T A
V W"
8 1 1 1
matches mismatches deletions insertions
Alignment":""2"x"k"matrix"("k"≥ m,"n")

SLIDE 2 “Goodness” of alignments
Given two sequences, there are many possible alignments ATTTTCCC ATTTACGC ATTT-TCCC ATTTA-CGC ATTTTCCC———————— ————————ATTTACGC
Edit distance: the total number of substitutions, insertions and deletions needed to transform one sequence to another
distance=2 distance=3 distance=16

SLIDE 3
Manhattan tourist problem
Imagine seeking a path (from source to sink) to travel (only eastward and southward) with the most number of attractions (*) in the Manhattan grid
Sink
* * * * * * * * * * *
Source
*

SLIDE 4 Recursive algorithm -> Dynamic programming
Function MT(n,m)
weight of the edge from (n-1,m) to (n,m)
weight of the edge from (n,m-1) to (n,m)
MT(x, y) returns the “most weighted” path from point (x, y) to the “sink”.

SLIDE 5 1 2 5 $5 1 $5 $5 3 3 5 3 3 5 10 $3 $5 $5 2 1 2 3 1 2 3
i source 1 3 8 5 8 8 4 9 13 8 12 9 15 9
1
16 S3,3$=/16
- Start from Sink.
- Find which of the two
edges gave the “max”. Take it.
How to find the optimal path

SLIDE 6 Recipe
- 1. Identify subproblems
- 2. Write down recursions
- 3. Make it dynamic-programming!

SLIDE 7 The edit distance problem
Match Insertion_X Insertion_Y
A-GCDEF AFGCDE-
A F G C D E A G C D E F

SLIDE 8
Minimum Edit Distance
For sequence X and Y

SLIDE 9 Optimal alignment
match match

SLIDE 10
Complexity

SLIDE 11
Is the edit distance the best way?
For sequence X and Y

SLIDE 12
Amino acids can share similar properties

SLIDE 13 Weighted edit distance
- To generalize scoring for DNA/RNA, consider a 4x4 scoring matrix
S.
- In the case of an amino acid sequence alignment, the scoring matrix
would be a 20x20 size.
- The addition of d is to include the score for comparison of a gap
character “-”.
- Two questions:
- (a) What should S be?
- (b) How do we find optimal scoring alignment?

SLIDE 14 Weighted edit distance
- To generalize scoring for DNA/RNA, consider a (4+1) x(4+1) scoring
matrix S.
- In the case of an amino acid sequence alignment, the scoring matrix
would be a (20+1)x(20+1) size.
- The addition of d is to include the score for comparison of a gap
character “-”.
- Two questions:
- (a) What should S be?
- (b) How do we find optimal scoring alignment?
Traditionally, people tend to maximize the alignment score with a negative gap penalty score

SLIDE 15
BLOcks SUbstitution Matrix (BLOSUM)
amino acids

SLIDE 16
BLOcks SUbstitution Matrix (BLOSUM)

SLIDE 17
Recursion for generalized edit distance
Complexity?

SLIDE 18
Gap score/penalty

SLIDE 19
Affine gap penalty
Question: How to develop an efficient dynamic programming algorithm for affine gap penalties?

SLIDE 20
Categories of pairwise alignments

SLIDE 21
Semi-global alignment

SLIDE 22
Semi-global alignment

SLIDE 23
- Long run time O(n4):
- In the grid of size n x n there are n2 vertices (i,j) that may serve as
a source.
- For each such vertex computing alignments from (i,j) to (i’,j’) takes
O(n2) time.
- This can be remedied by allowing every point to be the starting
point
Local alignment: naive algorithm

SLIDE 24
Local alignment: Smith-Waterman algorithm
Idea: start over from any entry!

SLIDE 25
Local alignment

SLIDE 26

SLIDE 27