Ensemble simulations with experimental restraints: Running at Scale
- or-
a designed interface for MD simulations
Peter Kasson, University of Virginia
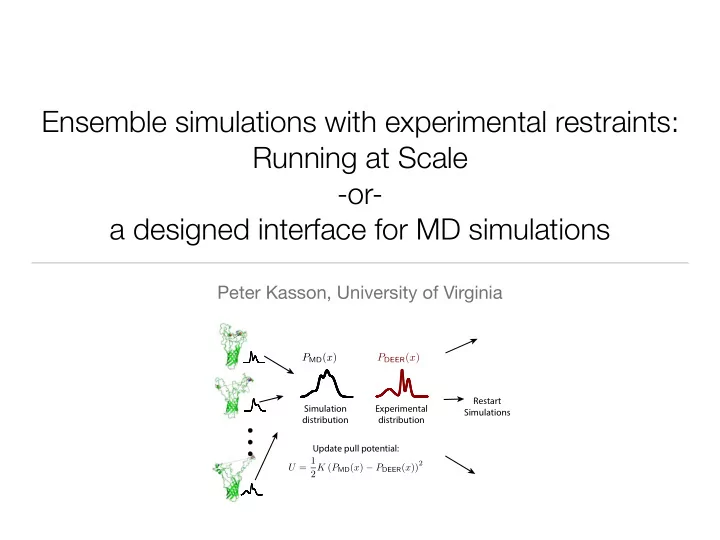
- Simulation
distribution Update pull potential: Experimental distribution Restart Simulations
Ensemble simulations with experimental restraints: Running at Scale - - PowerPoint PPT Presentation
Ensemble simulations with experimental restraints: Running at Scale -or- a designed interface for MD simulations Peter Kasson, University of Virginia Restart Simulation Experimental Simulations distribution distribution Update pull
distribution Update pull potential: Experimental distribution Restart Simulations
Image: deepideas.net
input structures run parameters simulation setup MD simulations trajectory analysis (colvars, clustering) sampling & setup for new simulations MD simulations trajectory analysis (colvars, clustering) ensemble results run inputs simulation trajectories clusters, collective variables run inputs
Execution manager
gmxapi.load_file params: [filename1, filename2, ...]
Data Input
gmxapi.md
MD Engine
>>> md = gmx.from_file([filename1, filename2, filename3, ...])
>>> gmx.run()
>>> md.add_dependancy(potential) >>> potential = myplugin.EnsembleRestraint(sites, *args, **kwargs) myplugin.mdmodule params: [...]
Plug-in module
gmxapi.ensemble_reduce params: [SUM]
Calculation on ensemble MD Engine Data Input MD Engine Data Input Calculation on ensemble Plug-in module
distribution Update pull potential: Experimental distribution Restart Simulations
Continue simulations
b) a)
Distance (nm) Probability Time (ns) J-S Divergence
MD Initial DEER
50 100 0.4 0.2 0.0
31-166 88-162 77-107 117-107
31 - 166 88 - 162 77 - 107 117 - 107