1
Molecular diagnostics of lymphoid malignancies
11th Course on Molecular Diagnostics Molecular Medicine Postgraduate School, Erasmus MC, November 3, 2017
- Dr. A.W. Langerak (Ton)
Laboratory for Medical Immunology, Dept. Immunology Erasmus MC, Rotterdam a.langerak@erasmusmc.nl
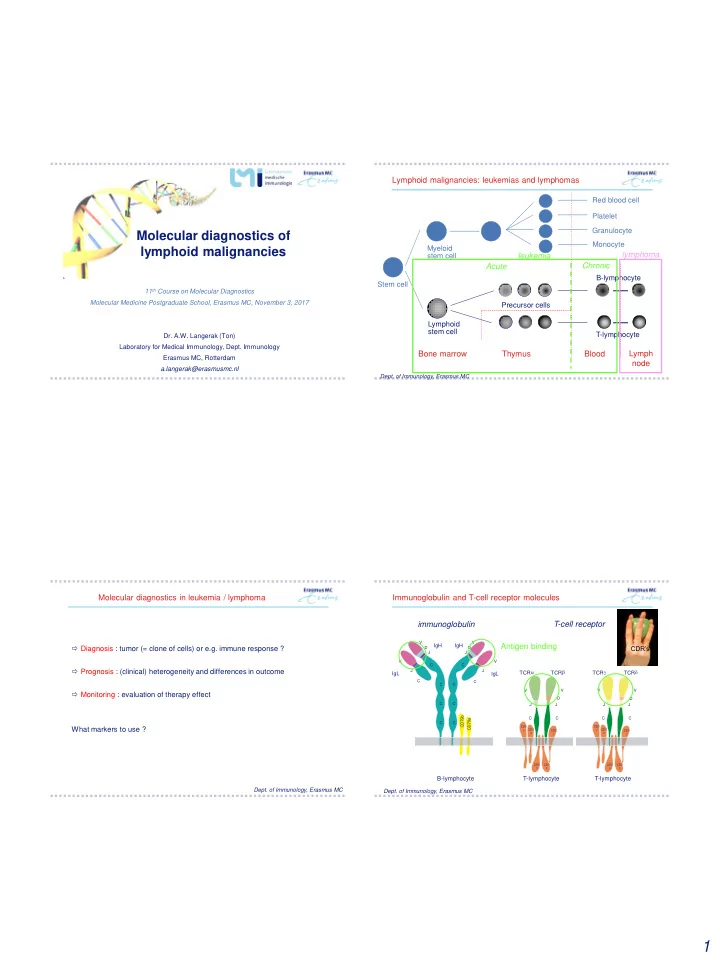
Stem cell Myeloid stem cell B-lymphocyte T-lymphocyte Precursor cells Red blood cell Platelet Granulocyte Monocyte
Bone marrow Thymus Blood
Lymphoid stem cell
Lymphoid malignancies: leukemias and lymphomas
- Dept. of Immunology, Erasmus MC
Lymph node lymphoma leukemia Acute Chronic
Diagnosis : tumor (= clone of cells) or e.g. immune response ? Prognosis : (clinical) heterogeneity and differences in outcome Monitoring : evaluation of therapy effect What markers to use ?
- Dept. of Immunology, Erasmus MC
Molecular diagnostics in leukemia / lymphoma
IgH IgL IgL
V V C C J J
IgH
V V D D J J C C C C C C C C C D 7 9 a C D 7 9 b
CD3 e CD3 d CD3 g CD3 x CD3 x
V J C V D J C
TCR a TCR b
B-lymphocyte T-lymphocyte
CD3 e CD3 d CD3 g CD3 x CD3 x
V J C V D J C
TCR g TCR d
T-lymphocyte
- Dept. of Immunology, Erasmus MC
immunoglobulin T-cell receptor Immunoglobulin and T-cell receptor molecules Antigen binding
CDR’s