- 1-
Workshop 7: (Generalized) Linear models
Murray Logan
July 19, 2017
Table of contents
1 Linear model Assumptions 1 2 Multiple (Genearalized) Linear Regression 15 3 Centering data 17 4 Assumptions 20 5 Multiple linear models in R 21 6 Model selection 26 7 Worked Examples 27 8 Anova Parameterization 29 9 Partitioning of variance (ANOVA) 35 10 Worked Examples 37
- 1. Linear model Assumptions
1.1. Assumptions
- Independence - unbiased, scale of treatment
- Normality - residuals
- Homogeneity of variance - residuals
- Linearity
1.2. Assumptions
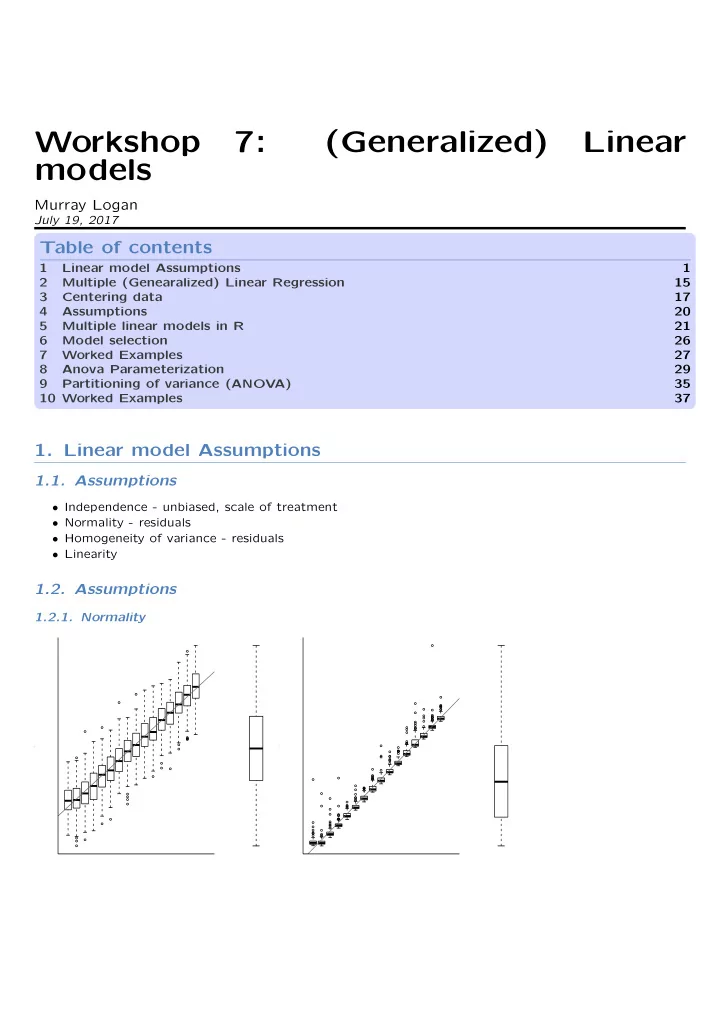
1.2.1. Normality
- y
- y