Gene regulation, protein networks and disease a computational - PowerPoint PPT Presentation
Gene regulation, protein networks and disease a computational perspective Ron Shamir School of Computer Science Tel Aviv University CPM Helsinki July 3 2012 1 1 Outline Finding regulatory motifs I, II, III Utilizing
Gene regulation, protein networks and disease – a computational perspective Ron Shamir School of Computer Science Tel Aviv University CPM Helsinki July 3 2012 1 1
Outline • Finding regulatory motifs I, II, III • Utilizing case-control expression profiles and networks I, II DEGAS • Chromosomal aberrations in cancer 2
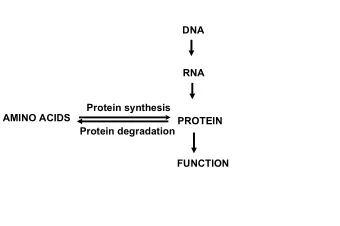
Regulation of Transcription • A gene’s ranscription regulation is mainly encoded in the DNA in a region called the promoter • Each promoter contains several short DNA subsequences, called binding sites (BSs) that are bound by specific proteins called transcription factors (TFs) TF TF 5 ’ 3 ’ Gene BS BS promoter
Position Weight Matrix (PWM) Score: product of 0.1 0.8 0 0.7 0.2 0 A base probabilities. 0 0.1 0.5 0.1 0.4 0.6 C Need score 0 0 0.5 0.1 0.4 0.1 G threshold for hits. 0.9 0.1 0 0.1 0 0.3 T ATGCAGGATACACCGATCGGTA 0.0605 GGAGTAGAGCAAGTCCCGTGA 0.0605 AAGACTCTACAATTATGGCGT 0.0151 4
C. Linhart, Y. Halperin Gen enome Res e Resea earch 08 08 I. Finding Regulatory Motifs 5
Motif discovery: The tw two-step tep s strategy egy Pr Promoter Co-reg Co egulated ed g gene set et sequences Cluster I Gene e exp xpression Clust stering microarray ays Cluster II Cluster III Motif discov overy Location a analysis (ChIP-chip, … (C …) Functional g l group (e (e.g., G GO term) 6
Amad adeus us A Motif Algorithm for Detecting Enrichment in mUltiple Species Supp pports d diverse m motif d disco covery t tasks: 1. Find ove over-re repre resented motifs in given sets of of genes. 2. Identify motifs with global s l spatial f l feature res given onl nly the genomic sequences. How? w? A general pipeli line a arc rchi hitecture for enumerating motifs. Different statistical sc scoring sc scheme mes of motifs for different motif discovery tasks. 7
Motif search algorithm Pipeline of refinement phases of increased complexity PW PWM Prepr process Mismat atch Merge Phases: Optimiza zation Cutoff = = 0.005 005 Mo Motif Mo Model el: k -mer List o of k- mers PW PWM 8
Scor coring ov over-rep epres esen ented ed m motifs Input put: Target set (size T ) = co-regulated genes Background (BG BG) set (size B ) = entire genome Mo Motif enri richment s sco cori ring: t B b Hyper-geom ometric T GC GC-conte tent Binne nned e enrichment nt s score 20-40 20 40% 40 40-60 60% B 1 B 2 0.4-0.7kb Bino nomi mial T 2 T 1 b 1 b 2 Length Le kbp B 3 B 4 0.7-1kbp T 4 b 4 T 3 b 3 bp 9
Metazoan motif discovery benchmark: 42 42 targ rget s sets of of 26 26 TFs, s, 8 8 miRNAs As from from 29 29 studies s (expre ression on, C , Chip-ChIP hIP,..) ,..) i in hu human, , mou ouse, , fly fly, w , worm orm. All ll m mot otifs fs a are re experi rimentally ve veri rified Ave verage t targ rget s set size: : 400 400 genes ( (383 383 Kb Kbp) ) 10
11
12
Amade deus s – Global spatial analysis Co-re Co regula lated g gene set Gene e expression on Location anal analysis ( (ChIP-chip, … …) Promoter microarrays sequences Functi tional g group ( p (e.g., G GO te term) m) Output 13 Motif(s)
Task II : Glo lobal a l analy lyse ses Scores for spatial features of motif occurrences In Input: Sequences (no target-set / expression data) Motif if s scorin ing: Localization w.r.t the TSS TSS SS 5’ Strand-bias Chromosomal preference 14
Global analysis: Chromosomal preference in C. elegans Input: t: Re Results: Novel m l motif on on chro rom IV IV All ll wo worm promoters rs (~ (~18 18,000 00) ) Score re: : chromosomal al prefere rence 15
Global analysis: Chromosomal preference in C. elegans Input: t: All ll wo worm promoters rs ( (~18 18,000 000) ) Score re: : chrom hromosomal p pre refe ference Re Results: Novel m l motif on on chr hrom om IV IV 16
Y. Halperin, C. Linhart, I. Ulitsky NAR AR 1 0 1 0 II. Finding Transcriptional Programs 17
Goal Given expression profiles, find the transcriptional programs active in them: - the co-regulated genes, - the motifs that govern their co- regulation
Our goal oal: b : bypas ass t the two-step a approac ach Co-regulated gene set Expression data Simultaneous s Promoter Cluster I Gene infer erence o e of the e sequences expression motif tifs a and the Clustering microarrays Cluster II exp pr p profiles o of their ir t targe gets ts Cluster III Output Motif(s) 19
Allegro: expression model Discretization of expression patterns Discrete e expression on Ex Expressi ssion p pattern Pattern ( (DEP EP) e 1 =Up (U) ≥ 1.0 e 2 =Same (S) (-1.0, 1.0) c 1 c 2 … c m c 1 c 2 … c m e 3 =Down (D) ≤ -1.0 g -2.3 -0.8 1.5 g D S … U Condition frequency matrix (CFM) c 1 c 2 … c m F = U 0.05 0.1 … 0.78 S 0.9 0.2 … 0.14 D 0.05 0.7 … 0.08 Condition weight matrix (CWM WM) f = ( W ) ( R= { r ij } is the BG CFM) ij F log r ij ⇒ Log-likelihood ratio (LLR LLR) score 20
Allegro overview 21
Yeast osmotic shock pathway ~6,000 genes, 133 conditions [O’Rourke et al. ’04] Allegro can discover multiple motifs with diverse expression patterns, even if the response is in a small fraction of the conditions Extant two-step techniques recovered only 4 of the above motifs: K-means/C /CLI LICK + + Amadeus/W /Weeder: RRPE, PAC, MBF, STRE 22 Iclust st + + FIRE: E: RRPE, PAC, Rap1, STRE
3’ ’ UT UTR R an anal alysis: Hu Human an st stem c cells s ~14,000 genes, 124 conditions (various types of proliferating cells) [Mueller et. al, Nature’08] Biases in length / GC-content of 3’ UTRs, e.g.: 100 highly-expressed genes in… 3’ UTR: length GC Embryoid bodies 584 47% Undifferentiated ESCs 774 44% ESC-derived fibroblasts 1240 39% Fetal NSCs 1422 43% ( ESCs = embryonic stem cells, NSCs = neural stem cells) Extant methods / Allegro with HG score: report only false positives 23
Hu Human an st stem cells: s: results using binned score miRN RNA targets s Current expressi ssion expressi ssion knowledge Most highly expressed miRNAs in human/mouse ESCs Abundant & functional in neural cell lineage Expressed specifically in neural lineage; active role in neurogenesis 24 miRNA expression from [Laurent ’08]
Yonit Halperin Chaim Linhart Igor Ulitsky Yaron Orenstein 25
Open questions Better PWM inference: new scores, algs Richer models for in vivo / in vitro data – really helpful or diminishing return? How to evaluate model quality: match to literature? Ranking based? In vivo? In vitro? Integration of motif finding & expression Principled means to find motif pairs 26
I. Ulitsky, R. M. Karp RECOMB 09 09 I. Ulitsky, A. Krishnamurthy, R. M. Karp PLo LoS One ne 1 0 1 0 Using expression profiles and protein networks to understand cancer I 27 27
DNA chips / Microarrays • Simultaneous measurement of expression levels of all genes. • Global view of cellular processes. • > 800,000 profiles available in ArrayExpress 28
Protein-protein interactions (PPIs) • A regulates/binds to B • High throughput: abundant, noisy • Large, readily available resource 29
Case/control studies • A typical study: 100s expression profiles of sick (case) & healthy samples (control) individuals genes • Classification: Given a partition of the samples into types, classify the types of new samples • Can the network help? sick healthy ? 30
The network angle • Integrate case-control profiles with network information • Extract dysregulated pathways specific to the cases • Account for heterogeneity among cases • Meaningful pathway: connected 31
Preprocessing • For each gene, use the Control 1 Control 2 Control 3 Control 4 Case 1 Case 2 Case 3 distribution of values among the controls to A B decide if the gene is C dysregulated in each of D E the cases Case 1 Case 2 Case 3 A Case 1 B 0 A 1 1 B 0 1 1 C Case 2 0 0 C 1 D 0 0 D 1 Case 3 1 E 1 1 E 32
Dysregulated pathway • Input: – Bipartite graph: genes, cases – Edge (gene g, case c) if g is dysregulated in c – A network over the genes • Dysregulated pathway (DP): smallest connected subnetwork s.t. A A sufficiently many genes are ≥k Case 1 Case 1 B B dysregulated in all but few cases ≤l C C Case 2 Case 2 • Small pathway focused disease D D Case 3 Case 3 explanation E E • Min connected set cover problem k= 2,l= 1 33
Complexity • Set cover problem: Given sets of elements, find fewest sets that cover all elements k l G Problem 1 0 Clique Set cover k 0 Clique Set k-cover 1 >0 Clique Partial set cover 1 0 Any Connected set cover (Shuai & Hu 06) • All are NP-Hard • Devised approximation and heuristic algs DysrEgulated Gene set Analysis via Subnetworks 34 DEGAS
Recommend
More recommend
Explore More Topics
Stay informed with curated content and fresh updates.