Graph estimation and selection strategies using the focused information criterion Eugen Pircalabelu
Joint work with Gerda Claeskens, L. Waldorp and S. Jahfari ORSTAT and Leuven Statistic Research Center FICology Workshop, Oslo, May 9-11th, 2016
1 / 24
FIC graphs
Goal: estimate cerebral pathways between brain regions (ROIs) Motivation: ROIs ‘collaborate’/connect to other ROIs: functional connectivity estimate and summarize the structure of the brain graphs targeted FIC approach to focus on parts of the brain How: Framework with both directed & undirected edges Lagged & contemporaneous ROI effects Penalized nodewise models with graph ‘composition’ rules Main contribution: select targeted (via the focus) graphs that account for ROI effects general framework where nodes can be other entities, not just brain ROIs
2 / 24
Motivation
fMRI data: 8 subjects in a resting state study i.e. subjects are not performing any task; parcellation of the brain was obtained containing 68 (further split to 448) regions of interest (ROIs); for each region, 240 volumes of the BOLD signal were obtained;
3 / 24
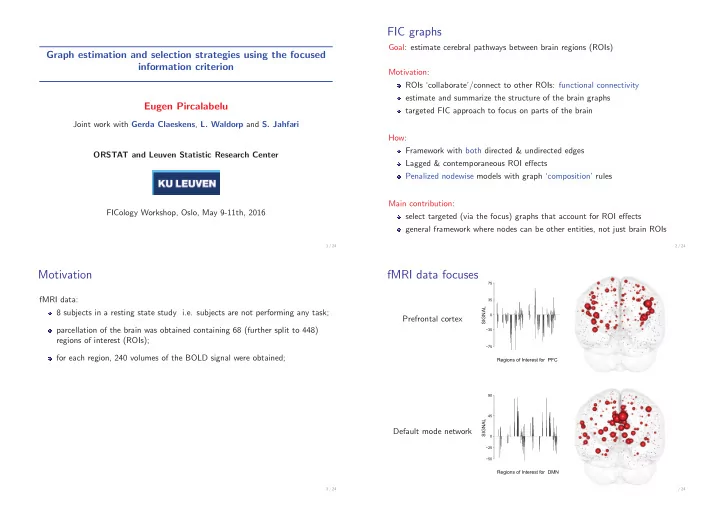
fMRI data focuses
Prefrontal cortex
SIGNAL
−75 −35 35 75
Regions of Interest for PFC
Default mode network
SIGNAL
−50 −25 45 90
Regions of Interest for DMN
4 / 24