CCS 2018 Satellite: DOOCN
Thessaloniki 27 September 2018
SIMPLICIAL COMPLEXES: EMERGENT HYPERBOLIC NETWORK GEOMETRY AND FRUSTRATED SYNCHRONIZATION Ginestra Bianconi
School of Mathema-cal Sciences, Queen Mary University of London, London, UK
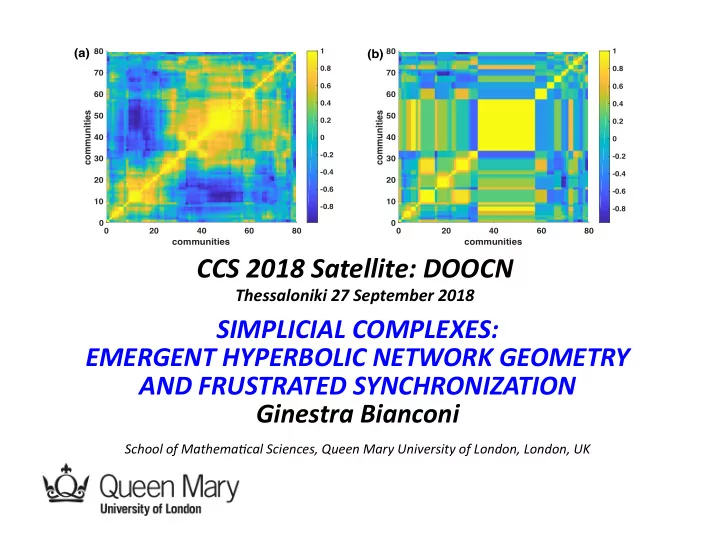
20 40 60 80
communities
10 20 30 40 50 60 70 80
communities
- 0.8
- 0.6
- 0.4
- 0.2
0.2 0.4 0.6 0.8 1
20 40 60 80
communities
10 20 30 40 50 60 70 80
communities
- 0.8
- 0.6
- 0.4
- 0.2
0.2 0.4 0.6 0.8 1
(a) (b)